Genetic Factors for Coronary Heart Disease and Their Mechanisms: A Meta-Analysis and Comprehensive Review of Common Variants from Genome-Wide Association Studies
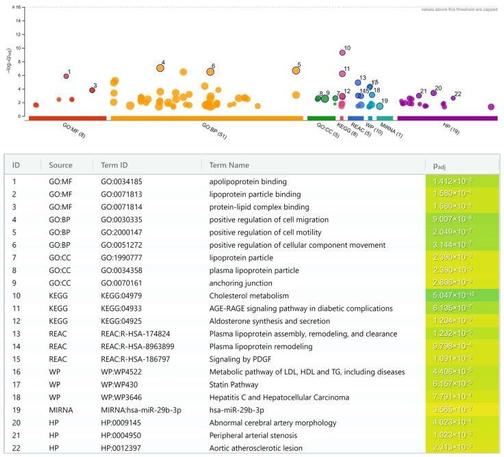
Genome-wide association studies (GWAS) have discovered 163 loci related to coronary heart disease (CHD). Most GWAS have emphasized pathways related to single-nucleotide polymorphisms (SNPs) that reached genome-wide significance in their reports, while identification of CHD pathways based on the combination of all published GWAS involving various ethnicities has yet to be performed. We conducted a systematic search for articles with comprehensive GWAS data in the GWAS Catalog and PubMed, followed by a meta-analysis of the top recurring SNPs from ≥2 different articles using random or fixed-effect models according to Cochran Q and I2 statistics, and pathway enrichment analysis. Meta-analyses showed significance for 265 of 309 recurring SNPs. Enrichment analysis returned 107 significant pathways, including lipoprotein and lipid metabolisms (rs7412, rs6511720, rs11591147, rs1412444, rs11172113, rs11057830, rs4299376), atherogenesis (rs7500448, rs6504218, rs3918226, rs7623687), shared cardiovascular pathways (rs72689147, rs1800449, rs7568458), diabetes-related pathways (rs200787930, rs12146487, rs6129767), hepatitis C virus infection/hepatocellular carcinoma (rs73045269/rs8108632, rs56062135, rs188378669, rs4845625, rs11838776), and miR-29b-3p pathways (rs116843064, rs11617955, rs146092501, rs11838776, rs73045269/rs8108632). In this meta-analysis, the identification of various genetic factors and their associated pathways associated with CHD denotes the complexity of the disease. This provides an opportunity for the future development of novel CHD genetic risk scores relevant to personalized and precision medicine.