Genetically Significant Hereditary Mutations Associated with Breast and Ovarian Cancer among Ukrainian Women - Cytology and Genetics
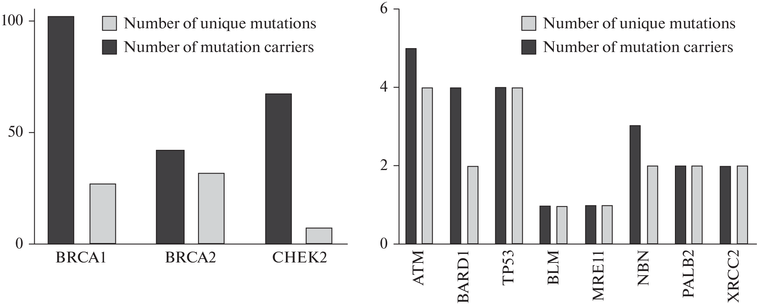
Abstract The study of mutations associated with the syndrome of hereditary breast cancer (BC) and ovarian cancer (OC) is extremely important for understanding the genetic risks among Ukrainian women. In this work, we conducted NGS study of mutations in the genes associated with the syndrome of hereditary BC and OC in 1090 women who had indications for testing and 407 women from the control group. As a result of the study, 233 patients with mutations were identified, including 84 unique variants. The largest number of mutations was registered in the BRCA1 (102), BRCA2 (42), and CHEK2 (67) genes. In the BRCA1 gene, the following mutations were the most frequent: c.5266dup (p.Gln1756fs), 45 cases; c.181T>G (p.Cys61Gly), 13; c.1510del (p.Arg504fs), 6; c.4035del (p.Glu1346fs), 5. The BRCA1 gene is a key risk factor for the development of cancer at a young age, while the CHEK2 is more frequently associated with cancer at an older age. The Population Attributable Risk (PAR) for the BRCA1 is 7.49%, which makes this gene a major risk factor. The odds ratio for mutations in the CHEK2 is 1.84 (95% CI: 1.01–3.34), while the PAR is 2.2%. The highest frequency of mutations (BRCA1 and BRCA2) was registered in the Central region of Ukraine. We for the first time performed a large-scale genetic study of the prevalence of hereditary mutations associated with the syndrome of hereditary BC and OC in Ukraine, the frequency of the most common mutations in the studied and control cohorts were determined.