🧬 Ready for AI to crack the RNA-disease code?
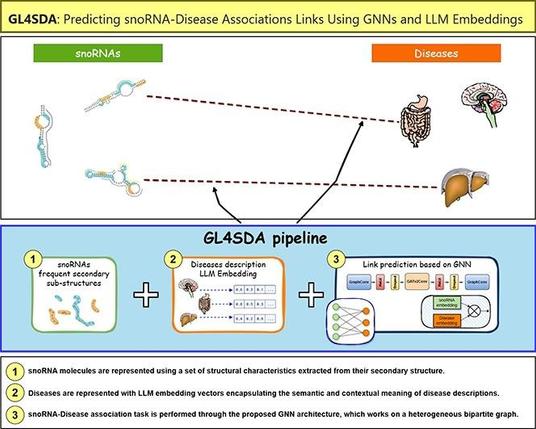
🔗 GL4SDA: Predicting snoRNA-disease associations using GNNs and LLM embeddings. Computational and Structural Biotechnology Journal, DOI: https://doi.org/10.1016/j.csbj.2025.03.014
📚 CSBJ: https://www.csbj.org/
#AIinBiomedicine #GraphNeuralNetworks #LLM #ncRNA #snoRNA #CancerResearch #Bioinformatics #PrecisionMedicine #XAI #Genomics