A Bioinformatician, Computer Scientist, and Geneticist lead bioinformatic tool development - which one is better? http://biorxiv.org/cgi/content/short/2024.08.25.609622v1?rss=1
Paul Gardner
- 108 Followers
- 129 Following
- 44 Posts
Bioinformatician. ex-Whāngārā. New Zealand. A/Prof. Overly fond of RNA.
What's the saying, don't believe a metric unless you've invented it yourself?
'Our work demonstrates that several widely used [gene prediction] tools are neither accurate nor computationally efficient for the protein-coding sequence detection problem. In fact, just three of nine tools significantly outperformed a naive scoring scheme.'
Flawed machine-learning confounds coding sequence annotation https://www.biorxiv.org/content/10.1101/2024.05.16.594598v1?med=mas
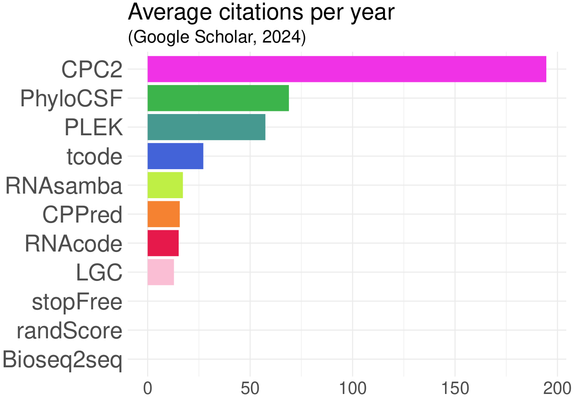
Finally, popularity turns out to be a terrible way to select a software tool. We have once again found that popular tools can be both inaccurate and slow on realistic, independent datasets. #citationmetrics 10/n
It is important to match confounding variables like homology, length & C+G in your +ve & -ve sets. Also, new genomes/transcriptomes have sequence & assembly error, this should be modelled too! #MachineLearning 9/n
We emphasize the importance of using realistic, diverse datasets for tool training and testing to avoid biases and improve tool reliability. #DataScience 8/n
Timing data showed that modern tools are not only often inaccurate but also computationally inefficient. Some widely used tools are both slow and inaccurate! #ComputationalBiology 7/n
We note a large discrepancy between self-reported tool accuracies and our independent evaluations, suggesting a need for more rigorous validation in the field. 6/n
The tools using evolutionary conservation (alignment inputs), like RNAcode and PhyloCSF, had much higher accuracy over single-sequence methods in identifying protein-coding regions. #EvolutionaryBiology 5/n
The results show that only three of nine popular tools significantly outperform a naive scoring scheme (longest stop-free subsequence). The underperforming tools are widely used by the lncRNA community. 4/n