For once I did not spend much time on biological networks (maybe inadvisable at a course whose subtitle contains the words "network modeling"), but instead presented SMuGLasso, a sparse multitask group lasso for GWAS in diverse populations that we developed with Asma Nouira and that was published just last week. https://journals.plos.org/ploscompbiol/article?id=10.1371/journal.pcbi.1012734
SMuGlasso groups SNPs by LD blocks (as done by Alia Dehman, Christophe Ambroise and Pierre Neuvial in 2015), and treats genetically homogeneous subpopulations of a heterogeneous data set as different but interlinked tasks. An additional sparsity constraints allows us to identify population-specific loci.
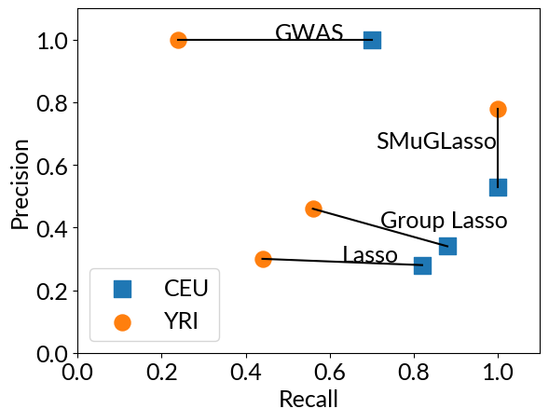
On simulations, we show that SMuGLasso has much better recall than classical GWAS (at the expense of more false positives, but fewer than other comparison partners).
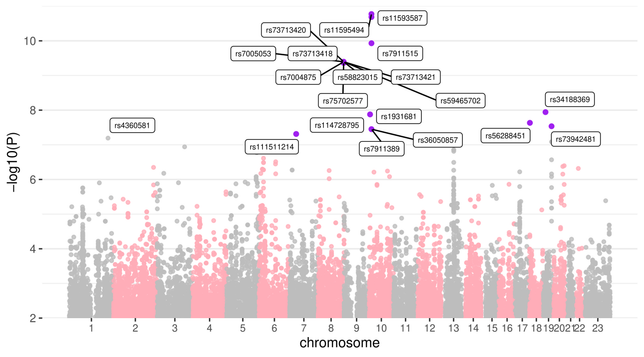
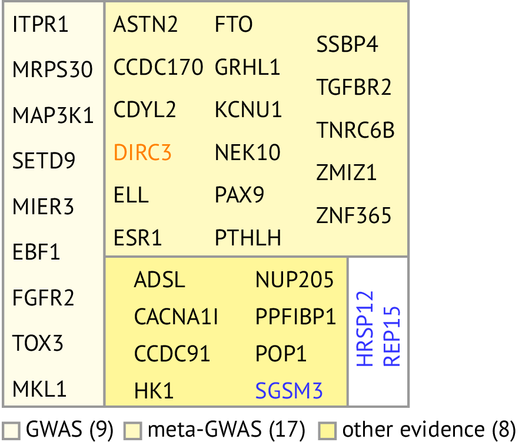
On real data, we observe that SMuGLasso recovers all the genes identified by a classical GWAS, as well as some that were identified by a meta-GWAS that included our dataset. All in all, only 2 of the 27 identified genes look likely to be false positives.
#machineLearning #genomics #GWAS