A resource of RNA-binding protein motifs across eukaryotes reveals evolutionary dynamics and gene-regulatory function.
Deciphering disordered regions controlling mRNA decay in high-throughput.
#IntrinsicallyDisorderedRegions #IDRs #mRNADecay #RNABindingProteins #RBPs #BuddingYeast
Improved RNA stability estimation through Bayesian modeling reveals most Salmonella transcripts have subminute half-lives.
Are you interested in RNA-binding proteins (#RBPs)? Then we have great news for you: our new EMBL Conference 'The expanding world of RBPs: from posttranscriptional control to riboregulation' has just opened for registrations! 🦠🧪
💻 https://s.embl.org/rbp25-01
📥 Abstract deadline: 8 December 2024
📆 11 – 14 March 2025
📍 EMBL Heidelberg and Virtual
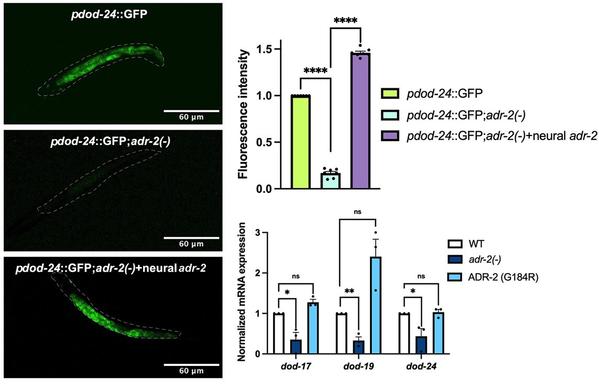
ADAR-mediated regulation of PQM-1 expression in neurons impacts gene expression throughout C. elegans and regulates survival from hypoxia
An animal’s capacity to thrive hinges on the nervous system translating environmental information into physiological responses. This study identifies a post-transcriptional mechanism whereby ADAR RNA binding proteins regulate the key transcriptional effector PQM-1 in the C. elegans nervous system to promote survival in low oxygen.
Some years ago, we developed the PTex method to purify RNA-protein complexes from cells and tissue using UV cross-linking and liquid phase separation. The method, along with similar ones, has been used to identify #RBPs cell-wide. However, finding proteins that bind to a single RNA, is much harder.
There is a very interesting follow-up now that uses a similar idea but targets individual #mRNA or #RNA species. Its still a preprint but it looks good imho:
Very pleased to see our paper is now in print:
Time-dependent regulation of cytokine production by RNA binding proteins defines T cell effector function https://www.cell.com/cell-reports/fulltext/S2211-1247%2823%2900430-8
here we show how RNA binding proteins regulate the magnitude and timing of cytokine expression in human T cells. and how the identified RBPs act a different time points and through different mechansism. The wonder world of RNA...
#RNA #RBPs #Immunology #Tcells #cytokines #regulation #proteomics
New preprint out where we apply gradient RNA sequencing and mass spec (Grad-seq) to Enterococcus faecalis and faecium to identify RNA-protein complexes #RBPs #microbiology #GradSeq #RNAseq #bacteria