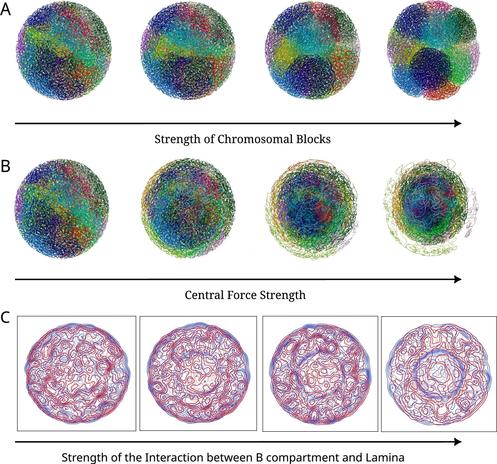
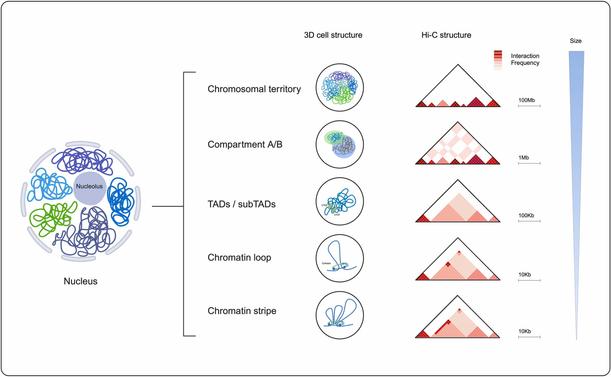
🧬 Ever wondered how the genome actually looks when it folds inside the nucleus — and how fast we can simulate it?
🔗 Multiscale molecular modeling of chromatin with MultiMM: From nucleosomes to the whole genome. Computational and Structural Biotechnology Journal, DOI: https://doi.org/10.1016/j.csbj.2024.09.025
📚 CSBJ: https://www.csbj.org/
#ChromatinModeling #3DGenome #ComputationalBiology #Genomics #MolecularModeling #GenomeArchitecture #ChromatinStructure #HiC #ATACSeq #Biophysics #StructuralBiology