Release Excited Squid Patch · nf-core/lsmquant
Fixed
PR#57 - Fix input declaration of the model file for the module: numorph3dunet to be reused for all inputs.
Remove conda batch from readme, the pipeline does not support conda.
PR#66 - Set ma...
Pipeline release! nf-core/rnafusion v4.1.3 - 4.1.3!
RNA-seq analysis pipeline for detection of gene-fusions
Please see the changelog: https://github.com/nf-core/rnafusion/releases/tag/4.1.3
#fusion #fusiongenes #genefusion #rna #rnaseq #nfcore #openscience #nextflow #bioinformatics
Release 4.1.3 · nf-core/rnafusion
What's Changed
Fusionreport Fixes (Cosmic etc pp) by @apeltzer in #815
make fasta input of star a value channel by @nvnieuwk in #816
Bump 4.1.3 by @apeltzer in #818
PR for 4.1.3 release by @apelt...
Pipeline release! nf-core/magmap v1.1.0 - nf-core/magmap v1.1.0!
Best-practice analysis pipeline for mapping reads to a (large) collections of genomes
Please see the changelog: https://github.com/nf-core/magmap/releases/tag/1.1.0
#nfcore #openscience #nextflow #bioinformatics

Release nf-core/magmap v1.1.0 · nf-core/magmap
What's Changed
Fix saving of samtools output by @erikrikarddaniel in #178
Fix/bakta gff compatibility by @Laurenz0908 in #181
Template 3.5.2, Nextflow lint and version topic channels by @erikrikar...
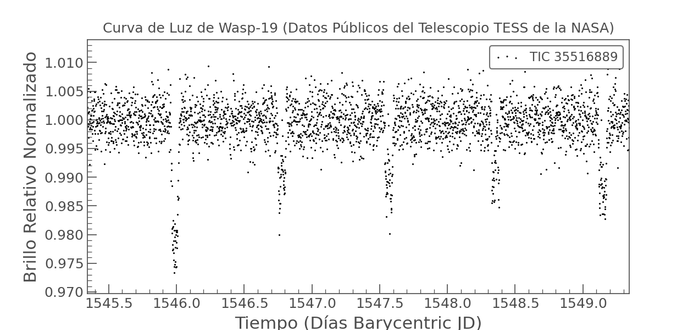
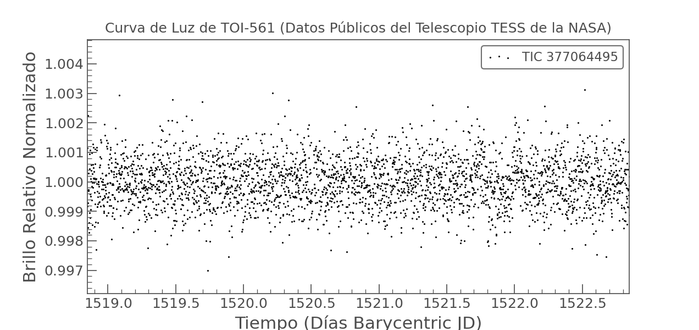
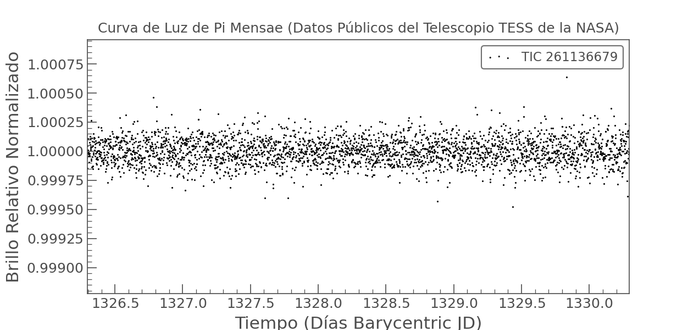
Aprovechando el fresquito para jugar a que
#Nextflow me busque exoplanetas usando datos de la NASA, ahí es nada
https://github.com/jagedn/search-planetasPipeline release! nf-core/pixelator v4.1.1 - nf-core/pixelator 4.1.1!
Pipeline to generate Proximity Network Assay data with Pixelator (Pixelgen Technologies AB)
Please see the changelog: https://github.com/nf-core/pixelator/releases/tag/4.1.1
#molecularpixelation #pixelator #pixelgentechnologies #proteins #singlecell #singlecellomics #nfcore #openscience #nextflow #bioinformatics

Release nf-core/pixelator 4.1.1 · nf-core/pixelator
[4.1.1] - 2026-05-29
Enhancements & fixes
Improve documentation by @Aratz #216
Update pixelator by @Aratz #217
Software dependencies
Dependency
Old version
New version
pixelator
0.27.1
0.27.2
Pipeline release! nf-core/pixelator v4.1.0 - nf-core/pixelator 4.1.0!
Pipeline to generate Proximity Network Assay data with Pixelator (Pixelgen Technologies AB)
Please see the changelog: https://github.com/nf-core/pixelator/releases/tag/4.1.0
#molecularpixelation #pixelator #pixelgentechnologies #proteins #singlecell #singlecellomics #nfcore #openscience #nextflow #bioinformatics
Release nf-core/pixelator 4.1.0 · nf-core/pixelator
[4.1.0] - 2026-05-28
Enhancements & fixes
Update pixelator container to 0.27.1 by @johandahlberg #212
Denoise graph before sample calling step by @Aratz #212
Add ACE and PLS denoising by @Aratz #2...
Pipeline release! nf-core/pixelator v4.0.0 - nf-core/pixelator 4.0.0!
Pipeline to generate Proximity Network Assay data with Pixelator (Pixelgen Technologies AB)
Please see the changelog: https://github.com/nf-core/pixelator/releases/tag/4.0.0
#molecularpixelation #pixelator #pixelgentechnologies #proteins #singlecell #singlecellomics #nfcore #openscience #nextflow #bioinformatics

Release nf-core/pixelator 4.0.0 · nf-core/pixelator
[4.0.0] - 2026-05-19
Added
Add Proxiome V2 workflow support, enabling sample pooling and processing of up to 8,000 cells per run by @Aratz #204, #205
Enhancements & fixes
Use nextflow strict syn...
Pipeline release! nf-core/pairgenomealign v2.2.3 - nf-core/pairgenomealign v2.2.3 – Reitou mikan!
Pairwise genome comparison pipeline using the LAST software to align a list of query genomes to a target genome, and plot the results
Please see the changelog: https://github.com/nf-core/pairgenomealign/releases/tag/2.2.3
#comparativegenomics #dotplot #genomics #last #pairwisealignment #synteny #wholegenomealignment #nfcore #openscience #nextflow #bioinformatics

Release nf-core/pairgenomealign v2.2.3 – Reitou mikan · nf-core/pairgenomealign
v2.2.3 "Reitou mikan" - [May 20th 2026]
Fixed
Conforms to nf-core template version 4.0.2 (#107).
Improve description of what the pipeline does and how (#108).
Dependencies
Dependency
Old versi...
Pipeline release! nf-core/mhcquant v3.2.0 - 3.2.0 - Solitude!
Identify and quantify MHC eluted peptides from mass spectrometry raw data
Please see the changelog: https://github.com/nf-core/mhcquant/releases/tag/3.2.0
#dda #immunopeptidomics #massspectrometry #mhc #openms #peptides #nfcore #openscience #nextflow #bioinformatics

Release 3.2.0 - Solitude · nf-core/mhcquant
Added
Added support for single run quantification #438
Added per-sample search parameter support via samplesheet with SearchPreset column #439
Added PRIDE ID and SDRF sheet support #445
Added ion ...
Pipeline release! nf-core/rnafusion v4.1.2 - 4.1.2!
RNA-seq analysis pipeline for detection of gene-fusions
Please see the changelog: https://github.com/nf-core/rnafusion/releases/tag/4.1.2
#fusion #fusiongenes #genefusion #rna #rnaseq #nfcore #openscience #nextflow #bioinformatics