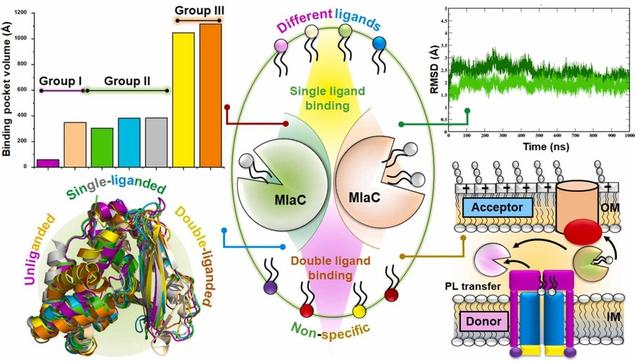
🧫 What secret molecular choreograph helps bacteria stay resilient against antibiotics and detergents?
🔗 Coordinated subdomain movements of MlaC regulate ligand binding and transport. Computational and Structural Biotechnology Journal, DOI: https://doi.org/10.1016/j.csbj.2025.05.031
📚 CSBJ: https://www.csbj.org/
#Microbiology #AntibioticResistance #MembraneBiology #StructuralBiology #ComputationalChemistry #MolecularDynamics #ComputationalBiology #Biophysics #ProteinStructure #DrugDiscovery #MolecularDocking

Identification of Bacterial FtsZ Effectors Targeting the Sites of Coumarin Binding - Cytology and Genetics
Abstract— There is a large group of bacterial FtsZ inhibitors, the biological activity of which has been confirmed biochemically. However, the sites of protein-ligand interaction for most of them remain unknown, significantly complicating the further search and combinatorial design of FtsZ inhibitors. This study presents the results of bioinformatic analysis of bacterial FtsZ effectors targeting sites of 4-hydroxycoumarin binding (BP1 and BP2). New data based on original results of pharmacophore screening, chemoinformatics, molecular docking, molecular dynamics simulations, AI-predictions, etc., are presented. The object of the study was a combined library of 379 compounds, formed based on revision of the structural database RCSB Protein Data Bank and biochemically proven FtsZ effectors from ChEMBL. Based on the results of a comprehensive study, 39 compounds were selected, of which 28 were identified as effectors of the sites BP1 and BP2, and another 11 as specific effectors of the site BP2 located in the BP2/IDC superpocket.
Architecting Tensor Core-Based Reductions for Irregular Molecular Docking Kernels
#CUDA #Chemistry #MolecularDocking #Package
https://hgpu.org/?p=30318

Architecting Tensor Core-Based Reductions for Irregular Molecular Docking Kernels
Tensor Cores (TCs) are specialized hardware units designed for efficient matrix multiplication and are widely utilized in deep learning workloads. However, their adoption in more irregular high-per…

Identification of FtsZ Protein Nucleotide-Binding Site Effectors Based on Cheminformatics and Structural-Biological Analysis - Cytology and Genetics
There is a large group of bacterial FtsZ inhibitors whose biological activity has been confirmed biochemically. However, the sites of protein–ligand interaction for most of them remain unknown, significantly complicating the further search and combinatorial design of FtsZ inhibitors. This study presents the results of bioinformatic analysis of bacterial FtsZ effectors, the interaction of which has been proven and documented in the ChEMBL database of biologically active molecules. With an integrated approach based on chemo- and bioinformatic methods and AI-based predictions, 23 inhibitors of nucleotide-binding site (NBS), as well as 16 new effectors of the interdomain cleft (IDC), were identified.
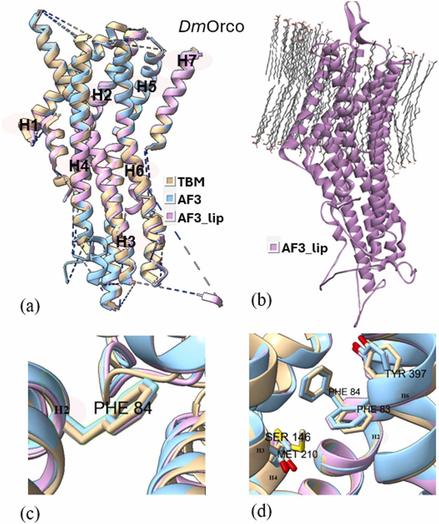
🦟 Could smarter protein modeling unlock eco-friendly pest control?
🔗 TBM preferred to AlphaFold 3 for functional models of insect odorant receptors. Computational and Structural Biotechnology Journal, DOI: https://doi.org/10.1016/j.csbj.2025.08.028
📚 CSBJ: https://www.csbj.org/
#ComputationalBiology #StructuralBiology #Bioinformatics #ProteinModeling #AlphaFold #MolecularDocking #DrugDiscovery #InsectScience #AIinBiology #TransmembraneProteins
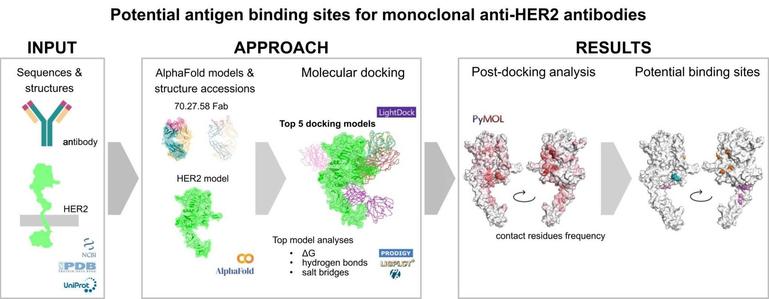
🔬 Can AI help decode how antibodies recognize cancer-related proteins like HER2?
🔗 Investigating and evaluating potential antigen binding sites for monoclonal anti-HER2 antibodies: The LightDock approach. Computational and Structural Biotechnology Journal, DOI: https://doi.org/10.1016/j.csbj.2025.06.001
📚 CSBJ: https://www.csbj.org/
#HER2 #CancerResearch #MonoclonalAntibodies #LightDock #AlphaFold #Bioinformatics #MolecularDocking #InSilicoBiology #ComputationalBiology #AntibodyResearch #StructuralBiology

Plant Histone Deacetylases: Their Classification and Inhibitor Search - Cytology and Genetics
Abstract Histone deacetylases is a family of enzymes pivotal in regulating numerous crucial cellular processes in both plant and animal cells. Plant histone deacetylases have been considerably less investigated in comparison to their human counterparts. This study aims to provide an in-depth characterization of histone deacetylases in two model plant proteins-species Arabidopsis thaliana and Oryza sativa. Phylogenetic analysis of their relationship to known human homologs has revealed their close relation to three classes of human histone deacetylases. Notably, the highest sequence homology among histone deacetylases from different evolutionary origins was observed between human HDAC6 and A. thaliana HDA5 (43.6% homology). Structural alignment results highlighted the conservation of catalytic domains and demonstrated a high activity of inhibitors against both histone deacetylases. Ligand-protein docking studies suggested a high potency of human histone deacetylase inhibitors against A. thaliana HDA5. These findings suggest the potential efficacy of human histone deacetylase inhibitors in modulating plant histone deacetylases, thereby enhancing growth regulation, development, and stress resistance in plants.

Identification of FtsZ Interdomain Cleft Effectors Based on Pharmacophore Search and Molecular Docking - Cytology and Genetics
Abstract There are a significant number of inhibitors of the bacterial FtsZ protein with biochemically confirmed biological activity, at the same time, their binding sites remain unclear. This significantly complicates further combinatorial design, and, in the current study, the results of a computational search for effectors of the Inter-Domain Cleft (IDC) site are presented. The actual research was based on the results of pharmacophore screening using the Pharmit service and molecular docking with CCDC GOLD and iGEMDOCK programs. The objective group was a combined library of 379 compounds, which was designed based on revision of the structural database of the RCSB Protein Data Bank and compounds from the ChEMBL database, for which direct interaction with FtsZ has been proven biochemically. According to the results of pharmacophore search, docking, and structural analysis, 88 effectors of the IDC site were identified. One more curcumin compound has been identified as a potential IDC site effector.