the third paper in the #iccs2025 collection is out: https://doi.org/10.1186/s13321-026-01203-8
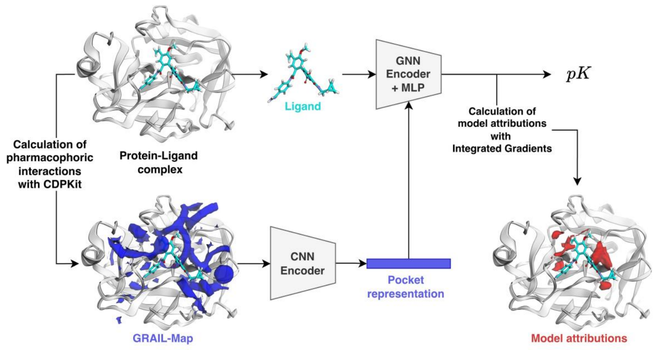
"GRIPHIN: grids of pharmacophore interaction fields for affinity prediction"
"We investigate whether a purely pharmacophoric representation of the protein pocket is sufficient to train a deep learning model for affinity prediction. For this purpose, we devise a hybrid model architecture from simple building blocks for affinity prediction."