📣 New Article from Network and Systems Medicine on #ScienceOpen!
🆕📄 'Identification of Transcriptional Regulators Using a Combined Disease Module Identification and Prize-Collecting Steiner Tree Approach' ➡️ https://drugrepocentral.scienceopen.com/hosted-document?doi=10.14293/NSM.25.1.0003
#REPO4EU #NetworkMedicine #Bioinformatics #TranscriptionFactors

Identification of Transcriptional Regulators Using a Combined Disease Module Identification and Prize-Collecting Steiner Tree Approach
<p xmlns:xsi="http://www.w3.org/2001/XMLSchema-instance" class="first" dir="auto" id="d4991e168">Transcription factors play important roles in maintaining normal biological function,
and their dysregulation can lead to the development of diseases. Identifying candidate
transcription factors involved in disease pathogenesis is thus an important task for
deriving mechanistic insights from gene expression data. We developed Transcriptional
Regulator Identification using Prize-collecting Steiner trees (TRIPS), a workflow
for identifying candidate transcriptional regulators from case–control expression
data. In the first step, TRIPS combines the results of differential expression analysis
with a disease module identification step to retrieve perturbed subnetworks comprising
an expanded gene list. TRIPS then solves a prize-collecting Steiner tree problem on
a gene regulatory network, thereby identifying candidate transcriptional modules and
transcription factors. We compare TRIPS to relevant methods using publicly available
disease datasets and show that the proposed workflow can recover known disease-associated
transcription factors with high precision. Network perturbation analyses demonstrate
the reliability of TRIPS results. We further evaluate TRIPS on Alzheimer’s disease,
diabetic kidney disease, and prostate cancer single-cell omics datasets. Overall,
TRIPS is a useful approach for prioritizing transcriptional mechanisms for further
downstream analyses.
</p>
Profound study on
#geneRegulation in E. coli:
#transcriptionFactors generally stabilize interaction with RNA polymerase and thus reduce expression range across promoters -> better maintenance of homeostasis.
https://www.science.org/doi/abs/10.1126/science.adv2064Rejuvenate Cells, Slow Aging - Jacob Kimmel and Dwarkesh Patel
#epigenome #transcriptionfactors
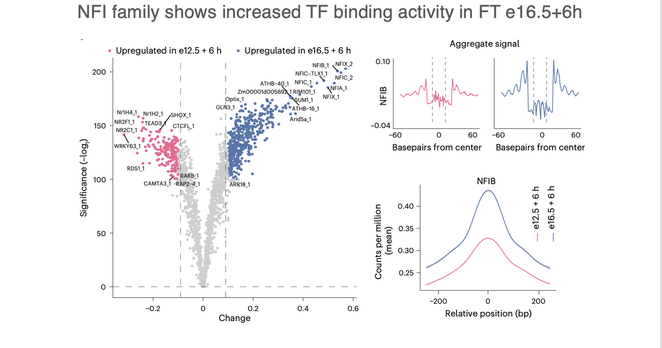
We also saw a shift in TF activity.
Footprinting showed increased NFI binding, especially NFIB, in late-born cohorts. These neurons activate maturation modules faster — possibly guided by chromatin remodeling during neurogenesis.
#NewsMayerlab #TFfootprints #TranscriptionFactors #scATACseq
(5/7)
Scleraxis (Genetics 🧬)
The scleraxis protein is a member of the basic helix-loop-helix superfamily of transcription factors. Currently two genes have been identified to code for identical scleraxis proteins.
https://en.wikipedia.org/wiki/Scleraxis
#Scleraxis #Genetics #GeneExpression #TranscriptionFactors
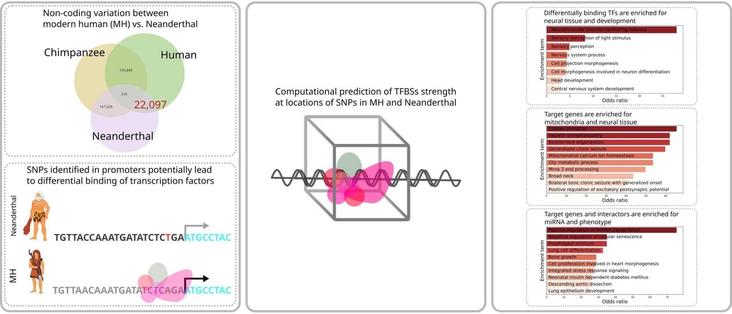
🧬 Why do humans think so differently from Neanderthals? The answer might lie in gene switches, not the genes themselves.
🔗 Evolution is in the details: Regulatory differences in modern human and Neanderthal. Computational and Structural Biotechnology Journal, DOI: https://doi.org/10.1016/j.csbj.2025.05.052
📚 CSBJ: https://www.csbj.org/
#Genomics #Neuroscience #Evolution #HumanOrigins #NeanderthalDNA #AutismResearch #HumanEvolution #GeneRegulation #Bioinformatics #ASD #RegulatoryGenomics #TranscriptionFactors

Elucidation of the Gene Regulatory Network Related to Spinal Muscular Atrophy - Cytology and Genetics
Abstract Spinal muscular atrophy (SMA) is an autosomal recessive heritable disorder leading to abnormalities and dysfunction of alpha motor neurons, paralysis, and eventual death due to respiratory failure. However, the gene regulatory mechanism related to SMA is still not completely clear. Here, we constructed the gene regulatory network of SMA, in which several SMA-related genes and transcription factors played important roles. In the process, 6544 differentially expressed genes (DEGs) associated with SMA were used and the enrichment analysis and gene regulatory network construction using machine learning was performed. The result showed that, firstly, p53 signaling and DNA replication are closely related to SMA. Then, there is a huge and complicated regulatory network guided by SMA, in which transcription factor SNAPC2, MZF1, and ZNF711 interacted closely with SMA-related genes (SMN1, SMN2) and played key roles in regulating genes of p53 signaling, DNA replication and other SMA-related GO (gene ontology) terms. The transcriptome data was well verified through real-time fluorescence quantitative PCR (RT-qPCR) using the peripheral blood of spinal muscular atrophy patients. Our study revealed the complicated gene regulation network of SMA, and uncover several important SMA-related genes, which it deepens our understanding of SMA-related regulatory mechanisms.

Transcription Factors in Cardiac Remodeling: Latest Advances - Cytology and Genetics
Transcription factors govern various functions in the cell such as proliferation, repair, regenerative programs, etc. Currently, there are no available drugs on the market or subjected to clinical trials that can combat cardiac remodeling. Understanding the role of transcription factors in cardiac remodeling may open the door to developing agents that can reverse structural remodeling of the heart and prevent heart failure. Recent studies shed light on the function of transcription factors involved in cardiac remodeling. Pharmacological modulation of signaling involving transcription factors may present as a novel mechanism for improving cardiac metabolism, promoting cardiac cell survival, etc. Various transcription factor-targeting agents were tested in animal models and showed promising results. Nevertheless, despite significant advances, the role of transcription factors in cardiac remodeling presents an extremely unexplored area. The main goal of this literature review was to summarize the latest advances made in our understanding of the role of transcription factors in cardiac remodeling and their potential to be used as molecular therapy targets.
Scientists recreate mouse stem cells f. gene older than animal life
https://www.sciencedaily.com/releases/2024/11/241118125716.htm
* unicellular Sox, POU transcription factors
* created mouse stem cells capable of gen. fully dev. mouse
* reshapes understanding of genetic/evol. origins of stem cells
* poss. critical for advent of stem cells, multicellularity
Emergence of Sox & POU transcript. factors predates origins of animal stem cells
https://www.nature.com/articles/s41467-024-54152-x
#StemCells #evolution #MolecularBiology #TranscriptionFactors #Sox #Pou

Scientists recreate mouse from gene older than animal life
An international team of researchers has achieved an unprecedented milestone: the creation of mouse stem cells capable of generating a fully developed mouse using genetic tools from a unicellular organism, with which we share a common ancestor that predates animals. This breakthrough reshapes our understanding of the genetic origins of stem cells, offering a new perspective on the evolutionary ties between animals and their ancient single-celled relatives.
Researchers challenge longstanding theories in cellular reprogramming
https://www.sciencedaily.com/releases/2024/11/241101123641.htm
* claim: neural crest stem cells = source of reprogrammed neurons found by other researchers
* refutes: any developed cell can be induced to switch identity to completely unrelated cell type thru infusion of transcription factors
Neural crest precursors f. skin are primary source of directly reprogrammed neurons
https://www.cell.com/stem-cell-reports/fulltext/S2213-6711(24)00291-1
#StemCells #MolecularBiology #CellBiology #TranscriptionFactors

Researchers challenge longstanding theories in cellular reprogramming
A team led by researchers has discovered that a group of cells located in the skin and other areas of the body, called neural crest stem cells, are the source of reprogrammed neurons found by other researchers. Their findings refute the popular theory in cellular reprogramming that any developed cell can be induced to switch its identity to a completely unrelated cell type through the infusion of transcription factors.