🚨 Application deadline extension! 🚨 Did you miss your chance to apply for #EMBLATAC? You're in luck, because now you have time until next Monday, 22 January, to do so! Apply here: https://s.embl.org/ata24-01
The aim of this course is to enable you to:
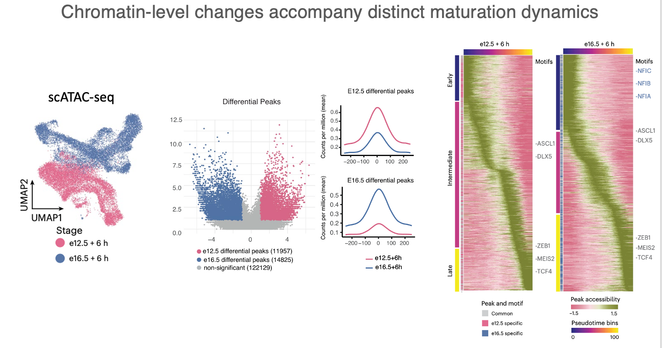
🔹Perform complete flow for single cell ATAC-seq experiments
🔹Process and quality control of single cell ATAC-seq data
🔹Visualise the data and call differences between samples
🔹Analyse and interpret the data using existing tools