RE: https://bsky.app/profile/did:plc:wf4t57jfj6xbvv4j7ebpy5z6/post/3mkntjym5r22f
RE: https://bsky.app/profile/did:plc:wf4t57jfj6xbvv4j7ebpy5z6/post/3mkntjym5r22f
Aidez à préserver l'efficacité des antibiotiques 💊 pour tous:
✅ Ne prenez que les antibiotiques qui vous ont été prescrits
✅ Demandez si cette prescription est le traitement recommandé de 1ère intention
✅ Lavez-vous les mains & faites-vous vacciner
Ministerstvo zdravotnictví 𝕏🔁 @[email protected]:
Vyhlašujeme výtvarnou soutěž pro děti Zastavme superbakterie! , určená pro základní školy. Děti se seznámí s problémem antibiotické rezistence a vytvoří obrázky, komiksy nebo příběhy o bakteriích.
Nejlepší díla odměníme a vystavíme v říjnu 2026!
🤖 로봇+AI 다 가졌다! LS티라유텍 주가 전망, 지금이라도 타야 하는 이유 (수익률 20% 공략법) ✅
LS티라유텍(322180)은 스마트팩토리 솔루션 및 자율주행 로봇(AMR) 분야의 선도 기업으로서, 최근 LS그룹으로의 편입과 하이테크 산업군의 자동화 수요 증가에 힘입어 시장의 뜨거운 관심을 받고 있습니다
2026년 5월 8일 기준, 최근 10거래일간의 주가 흐름은 단순한 기술적 반등을 넘어선 유의미한 시그널을 보여주고 있으며, 최신 데이터와 시장 동향을 정밀 분석하여, 투자 판단에 실질적인 도움이 될 수 있는 자료 공유,
LS티라유텍 일봉 차트 [자료:네이버]1. LS티라유텍 최근 주가 상승 요인 분석
최근 10거래일 동안 LS티라유텍의 주가는 견조한 우상향 곡선을 그리며 시장의 주도주로 부각되었으며, 이 상승의 핵심 동력은 다음과 같이 요약됩니다
- LS그룹 시너지 가속화: LS그룹 편입 이후 그룹 내 제조 계열사들의 스마트팩토리 고도화 프로젝트가 본격화되면서 캡티브(Captive) 물량 확보에 대한 기대감이 주가를 견인했습니다
- 특히 전력 인프라 및 이차전지 소재 부문과의 협업이 가시화되었습니다
- 하이테크 산업의 무인화 수요 폭증: 반도체와 이차전지 산업의 미세 공정화 및 인건비 상승에 대응하기 위한 무인 자동화 공장 시스템 수요가 급증하며, 동사의 통합 솔루션 역량이 부각되었습니다
- 기술적 변곡점 돌파: 장기 하락 추세선을 돌파하며 주요 이동평균선이 정배열 구간으로 진입함에 따라 기술적 매수세가 강하게 유입되었습니다
- 특히 5월 8일 발생한 VI(변동성 완화장치) 발동은 강력한 매수 에너지의 방출로 해석됩니다
- 외국인 및 기관의 수급 개선: 개인 위주의 거래에서 탈피하여, 기관 투자자들의 순매수가 유입되기 시작하며 수급의 질적 개선이 이루어졌습니다
2. 최근 호재 뉴스 요약
시장 현장에서 전해진 주요 소식들은 동사의 펀더멘털 강화와 직접적으로 연결되어 있습니다
- 스마트팩토리 통합 제어 소프트웨어 고도화: AI 기반의 예지 보전 기능이 강화된 새로운 버전의 소프트웨어가 출시되어 시장 점유율 확대를 예고했습니다
- 글로벌 공급망 확대 뉴스: 북미 지역의 이차전지 공장을 대상으로 한 물류 로봇 및 자동화 시스템 공급 논의가 구체화되고 있다는 소식이 전해지며 글로벌 성장성이 재조명되었습니다
- 정부의 ‘AI 제조 혁신’ 정책 수혜: 정부가 주도하는 중소·중견기업 대상 스마트공장 보급 확산 사업에서 동사의 솔루션이 우수 사례로 선정되며 정책적 수혜 기대감이 반영되었습니다
3. 최근 신용거래 비중과 잔고 동향 분석
신용잔고는 투자자들의 단기 향방을 가늠하는 척도입니다
- 신용 비중의 안정화: 주가 상승에도 불구하고 신용거래 비중은 급격히 늘지 않고 적정 수준을 유지하고 있습니다. 이는 과열 양상보다는 실수요 기반의 매수세가 유입되고 있음을 시사합니다
- 잔고 동향의 질적 변화: 과거 단기 차익을 노린 고레버리지 물량들이 상당 부분 소화되었으며, 현재는 주가 상승을 확신하는 중기 성향의 투자자들이 잔고의 주축을 이루고 있는 것으로 분석됩니다
- 반대매매 리스크 감소: 주가가 지지선을 확보하며 상승함에 따라 담보 부족으로 인한 강제 청산 리스크가 현저히 낮아졌습니다
4. 최근 공매도 비중과 동향 분석
코스닥 중소형주인 동사에게 공매도 동향은 주가 상방을 제한하는 주요 변수입니다
- 공매도 세력의 관망세: 최근 10거래일간 공매도 거래 대금 비중은 유의미하게 감소했습니다
- 주가의 하방 압력이 약해졌음을 의미하며, 이는 숏커버링(Short Covering) 유입 가능성을 높이는 요소입니다
- 대차잔고의 완만한 감소: 주식을 빌려 파는 행위가 줄어들면서 대차잔고가 완만하게 줄어들고 있습니다
- 세력들이 현재 주가 수준에서 추가 하락에 배팅하기보다는 상승 가능성에 무게를 두고 있음을 보여줍니다
5. 최근 시장심리와 리스크 요인 분석
심리적 지표와 잠재적 위험 요소를 동시에 고려해야 합니다
- 시장 심리(Sentiment): 극도의 공포 단계에서 벗어나 ‘탐욕’과 ‘낙관’ 사이의 중립 단계로 진입했습니다
- 투자자들 사이에서 “LS그룹의 진정한 수혜주”라는 인식이 퍼지며 매수 대기 자금이 풍부해진 상황입니다
- 리스크 요인 1 (실적 변동성): 수주 기반 산업 특성상 분기별 실적 편차가 클 수 있습니다
- 특히 적자 상태인 순이익 구조의 개선 속도가 시장 기대치에 못 미칠 경우 변동성이 커질 수 있습니다
- 리스크 요인 2 (거시 경제): 고금리 기조가 유지될 경우 설비 투자(CAPEX) 심리가 위축될 수 있으며, 이는 스마트팩토리 발주 지연으로 이어질 우려가 있습니다
6. 향후 주가 상승 지속가능성 분석
상승세가 “반짝 반등”에 그칠지, “대세 상승”의 서막일지에 대한 분석입니다
- 수주 잔고의 증가세: 현재 동사의 수주 잔고는 역대 최고 수준을 경신 중인 것으로 파악됩니다
- 이는 향후 1~2년간의 실적 가시성을 높여주며 상승 지속성을 뒷받침합니다.
- 이익률 개선 가시화: 저가 수주 물량이 해소되고 고부가가치 솔루션 비중이 높아지며 영업이익률이 턴어라운드하는 구간에 진입했습니다
- 숫자로 증명되는 상승세는 힘이 강합니다
- 업종 내 밸류에이션 재평가: 로봇 및 AI 테마 내에서도 실체가 있는 실적주로서의 평가를 받기 시작하며 멀티플(Multiple) 상향 조정이 이루어지고 있습니다
7. 향후 주목해야 할 이유 분석
투자자들이 LS티라유텍을 포트폴리오에 담아야 할 핵심 논거입니다
- 로봇 전문 자회사 ‘티라로보틱스’의 성장: 자회사인 티라로보틱스의 자율주행 물류로봇(AMR)이 북미 시장 등에서 기술력을 인정받고 있으며, 이는 모회사인 동사의 기업가치 증대로 직결됩니다
- LS그룹 내 ‘디지털 전환(DX)’ 컨트롤 타워: 단순 계열사를 넘어 그룹 전체의 스마트 제조 혁신을 주도하는 핵심 조직으로서의 위상이 강화되고 있습니다
- 산업용 소프트웨어의 국산화: 외산 소프트웨어가 장악했던 MES, 원격 제어 시장에서 국산화 점유율을 높이며 독보적인 해자(Moat)를 구축하고 있습니다
8. 향후 투자 적합성 판단
- 적합 대상: 하이테크 산업의 성장성을 믿는 중장기 투자자, LS그룹의 제조 경쟁력 강화에 베팅하는 투자자, 로봇 및 AI 자동화 테마의 실적주를 찾는 투자자에게 매우 적합합니다.
- 부적합 대상: 단기적인 급등락만을 쫓는 초단기 스캘퍼나, 실적 턴어라운드 과정의 변동성을 견디기 어려운 안정 지향형 투자자에게는 다소 공격적인 종목일 수 있습니다.
9. LS티라유텍 주가 전망과 투자 전략
종합적인 판단을 바탕으로 향후 주가 전망과 구체적인 대응 전략을 제시,
[주가 전망]
- 단기적 관점: 5월 8일의 VI 발동과 대량 거래 수반은 상방 매물을 상당 부분 소화했다는 증거
- 1차 목표가인 11,000원 선까지의 추가 상승 시도가 예상
- 중장기적 관점: 실적 개선세와 그룹사 시너지가 본격화되는 2026년 하반기에는 과거 전고점 부근인 13,000원 이상으로의 밸류에이션 점프가 충분히 가능할 것으로 전망
[투자 전략]
LS티라유텍은 단순한 테마주를 넘어 실질적인 ‘제조 혁신’의 아이콘으로 거듭나고 있습니다. 시장의 변동성에 일희일비하기보다는 동사가 가진 기술력과 그룹사 내 입지, 그리고 하이테크 산업의 성장에 초점을 맞춘다면 좋은 성과를 거두실 수 있을 것입니다
LS티라유텍(322180) 핵심 요약
LS티라유텍(322180)의 분석 내용을 핵심 중심으로 요약,
1. 주가 상승의 핵심 동력 (최근 10거래일)
- LS그룹 시너지: 그룹 편입 이후 계열사(이차전지, 전력 인프라) 대상 스마트팩토리 수주 가시화
- 기술적 전환점: 장기 하락 추세를 벗어나 대량 거래를 동반한 골든크로스 발생 및 VI 발동
- 수급 개선: 개인 중심에서 기관 및 외국인의 순매수세로 수급 주체가 전환되며 상승 탄력 강화
2. 시장 환경 및 내부 지표
- 호재 뉴스: AI 기반 스마트팩토리 소프트웨어 고도화 및 북미 이차전지 공장 대상 물류 로봇 공급 협의
- 건전한 지표: 주가 상승에도 불구하고 신용잔고 비중이 안정적이며, 공매도 비중은 감소하여 숏커버링(공매도 되사기) 기대감 상승
- 리스크 관리: 고금리에 따른 기업들의 설비투자(CAPEX) 지연 가능성 및 분기별 실적 변동성은 주의 요망
3. 향후 전망 및 투자 등급
- 주목 이유: 자회사 ‘티라로보틱스’의 자율주행 로봇(AMR) 성장성과 LS그룹의 디지털 전환(DX) 컨트롤 타워 역할 수행
- 상승 지속성: 역대 최고 수준의 수주 잔고와 이익률 개선(턴어라운드) 구간 진입으로 인해 우상향 기조 유지 전망
- 투자 적합성: 성장주와 실적주 성격을 동시에 갖추어, 로봇/AI 자동화 섹터 내 중장기 투자에 매우 적합
4. 투자 전략 가이드
- 목표가: 단기 11,000원 돌파 시도 후, 실적 확인 시 중장기 13,000원 이상 기대
- 매수 구간: 9,500원 ~ 9,800원 부근 분할 매수 (추격 매수 지양)
- 손절/지지선: 주요 지지선인 8,800원 이탈 시 비중 축소 권고
핵심 결론 : LS티라유텍은 단순 테마를 넘어 실질적인 수주와 그룹사 배경을 갖춘 종목입니다
단기 급등에 따른 눌림목을 활용해 비중을 확보해 나가는 전략이 유효해 보입니다.
(* 본 글은 정보 제공 목적이며, 투자 권유가 아닙니다. 투자 결정은 본인 책임하에 신중히 하시기 바랍니다)
단순 반등 아닌 찐 상승! 후성(093370) 향후 투자 전략.. 이 숫자만 기억하세요! 🧨에코프로머티, 또 한 번의 2배 가능? 전구체·탈중국 모멘텀 완전 해부글
- 5월 12, 2026
- 5월 11, 2026
- 5월 11, 2026
- 5월 11, 2026
- 5월 11, 2026
- 5월 11, 2026
🧫 Over-prescription of antibiotics is a major contributor to the rise in #AMR. IMI project VALUE-Dx investigated whether point-of-care (POC) testing could reduce antibiotic prescription.
💊The result? Having POC tests available did not lead to a reduction in prescriptions, and both clinician and patient perspectives play an important role in antibiotic prescriptions.
👉 https://link.europa.eu/DDgnw7
#IHITransformingHealth #HorizonEU #health #research #AntimicrobialResistance
💊 Die WHO stuft Antibiotikaresistenz #AMR als eine der 10 größten globalen Bedrohungen ein.
Wie ist die AMR-Lage in Deutschland? RKI-Forschende haben Daten aus 104 Laboren, 750 Allgemeinkrankenhäusern und 30.000 Praxen ausgewertet.
Die Analyse im #Bundesgesundheitsblatt
🔗 https://link.springer.com/article/10.1007/s00103-026-04234-6
🦠 Could we outsmart antibiotic resistance by turning bacteria against themselves?
🔗 Rational Targeting and gRNA Design for Enhancing Quorum Quenching in Pseudomonas aeruginosa PAO1. Computational and Structural Biotechnology Journal (CSBJ). DOI: https://doi.org/10.34133/csbj.0089
📚 CSBJ - A Science Partner Journal: https://spj.science.org/journal/csbj
#AntimicrobialResistance #AMR #CRISPR #GeneEditing #SyntheticBiology #SystemsBiology #Microbiology #Biotechnology #DrugResistance #QuorumSensing #Biofilms #Genomics
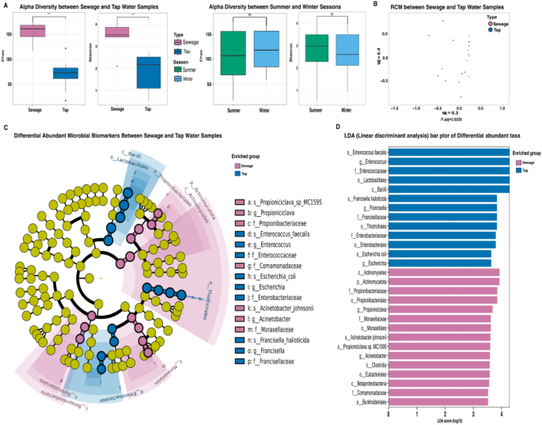
Metagenomic profiling of microbial communities and the resistome within Egyptian hospital wastewater and tap water - Scientific Reports
Antimicrobial resistance (AMR) is a worldwide health concern that compromises the successful treatment of a growing array of infectious diseases, particularly in low- and middle-income countries. AMR is exaggerated by the spread of antimicrobial resistance genes (ARGs) across humans, animals, and environmental reservoirs like water and soil. Hospital wastewater (HWW) is the main source of antimicrobial resistance in the environment. The current study used high throughput metagenomic nanopore sequencing to investigate the microbial abundance and ARGs associated with both HWW and tap water in five different hospitals in Cairo, Egypt. The bacterial community composition of the HWW microbiome identified 25 taxonomic families. The most abundant genera in HWW were Acinetobacter (6%) and Propioniciclav (5%) out of 101 unique genera while, the most abundant in tap water were Enterococcus (53%), Escherichia (15%), and Francisella (14%) out of 89 unique genera. Alpha diversity analysis revealed significantly greater microbial diversity in the HWW samples than in the tap water samples (P value > 0.05), moreover beta diversity analysis revealed a significant difference in the microbial community composition between the tap water and HWW samples (P value > 0.05) using Chao metric for richness estimation and Shannon metric for richness and evenness estimation. Total ARG analysis revealed absence of ARGs in tap water using the three databases, while comparable levels of ARGs were detected in HWW across the five hospitals. In total, 45, 28, and 28 ARG subtypes were identified in the HWW samples using ResFinder, CARD, and the NCBI AMRFinderPlus databases, respectively. The most abundant AMR mechanisms among the five hospitals were linked to the inhibition of protein synthesis. Using the ResFinder database, streptogramin resistance genes were most prevalent in Hospitals 1 and 5 (15% and 40%, respectively); using CARD, aminoglycoside, lincosamide, and macrolide resistance genes were most predominant (relative abundances 35–60%). Using NCBI AMRFinderPlus, streptomycin, tetracycline, and macrolide resistance genes were most prevalent (relative abundances 30.1–60%). Detection of plasmid replicons in HWW identified 39 different plasmid-associated replication genes via the PlasmidFinder database. The Col440l-1, colRNAI-1 and Col440ll-1 plasmid replicons were the most detected across the five hospitals with relative abundances of 16.6%, 10.9% and 9.6%, respectively. This study revealed different microbial communities among HWW and tap water in addition to the widespread occurrence of ARGs and AMR encoding plasmid replicons in the HWW in the five different hospitals in Cairo, Egypt indicating a significant risk associated with HWW, necessitating the implementation of preventative measures to avert their environmental diffusion. To our knowledge, this is one of the first Egyptian studies to apply Oxford Nanopore long-read metagenomic sequencing for simultaneous profiling of microbial communities and the resistome in HWW and tap water, using three ARG databases across five hospitals in two seasons.

Metagenomic profiling of microbial communities and the resistome within Egyptian hospital wastewater and tap water - Scientific Reports
Antimicrobial resistance (AMR) is a worldwide health concern that compromises the successful treatment of a growing array of infectious diseases, particularly in low- and middle-income countries. AMR is exaggerated by the spread of antimicrobial resistance genes (ARGs) across humans, animals, and environmental reservoirs like water and soil. Hospital wastewater (HWW) is the main source of antimicrobial resistance in the environment. The current study used high throughput metagenomic nanopore sequencing to investigate the microbial abundance and ARGs associated with both HWW and tap water in five different hospitals in Cairo, Egypt. The bacterial community composition of the HWW microbiome identified 25 taxonomic families. The most abundant genera in HWW were Acinetobacter (6%) and Propioniciclav (5%) out of 101 unique genera while, the most abundant in tap water were Enterococcus (53%), Escherichia (15%), and Francisella (14%) out of 89 unique genera. Alpha diversity analysis revealed significantly greater microbial diversity in the HWW samples than in the tap water samples (P value > 0.05), moreover beta diversity analysis revealed a significant difference in the microbial community composition between the tap water and HWW samples (P value > 0.05) using Chao metric for richness estimation and Shannon metric for richness and evenness estimation. Total ARG analysis revealed absence of ARGs in tap water using the three databases, while comparable levels of ARGs were detected in HWW across the five hospitals. In total, 45, 28, and 28 ARG subtypes were identified in the HWW samples using ResFinder, CARD, and the NCBI AMRFinderPlus databases, respectively. The most abundant AMR mechanisms among the five hospitals were linked to the inhibition of protein synthesis. Using the ResFinder database, streptogramin resistance genes were most prevalent in Hospitals 1 and 5 (15% and 40%, respectively); using CARD, aminoglycoside, lincosamide, and macrolide resistance genes were most predominant (relative abundances 35–60%). Using NCBI AMRFinderPlus, streptomycin, tetracycline, and macrolide resistance genes were most prevalent (relative abundances 30.1–60%). Detection of plasmid replicons in HWW identified 39 different plasmid-associated replication genes via the PlasmidFinder database. The Col440l-1, colRNAI-1 and Col440ll-1 plasmid replicons were the most detected across the five hospitals with relative abundances of 16.6%, 10.9% and 9.6%, respectively. This study revealed different microbial communities among HWW and tap water in addition to the widespread occurrence of ARGs and AMR encoding plasmid replicons in the HWW in the five different hospitals in Cairo, Egypt indicating a significant risk associated with HWW, necessitating the implementation of preventative measures to avert their environmental diffusion. To our knowledge, this is one of the first Egyptian studies to apply Oxford Nanopore long-read metagenomic sequencing for simultaneous profiling of microbial communities and the resistome in HWW and tap water, using three ARG databases across five hospitals in two seasons.
C. difficile S-layer’s role in...









