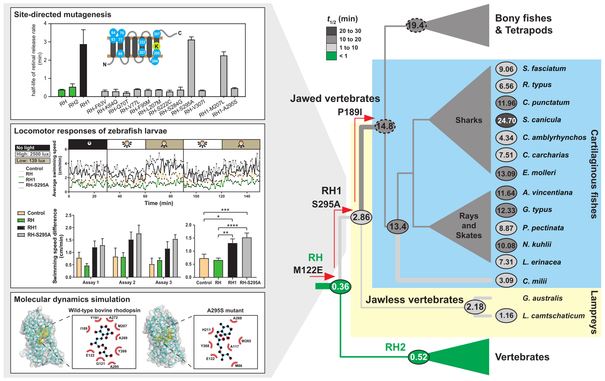
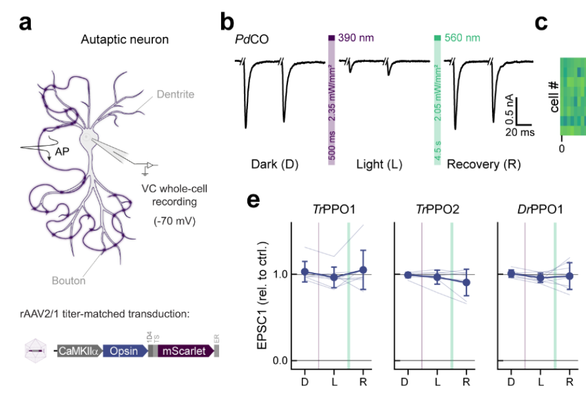
Cui et al. investigate the evolutionary trajectory leading to dim-light vision in vertebrates, concluding that it evolved via three main stages of molecular adaptation, at the origins of ancestral rhodopsin pigments.
Zhongmou Wins U.S. FDA Clearan...
#Zhongmou wins U.S. #FDA clearance for China's first optogenetic human trials https://www.linkedin.com/pulse/zhongmou-wins-us-fda-clearance-chinas-first-optogenetic-k1vte/ using ZM-02 for treating advanced retinitis pigmentosa; #RP #blindness #opsin #optogenetics #PRISM
"By bypassing damaged rod and cones, this mutation-agonistic approach is being developed for advanced retinal diseases, including retinitis pigmentosa and geographic atrophy (GA) in age-related macular degeneration."
We thank our postdoc Natalie Roberts for leading a stimulating lab-meeting today on #colour #vision and #opsin genes in damselflies and other insects for members of #SvenssonLab and #TsuboiLab.
In the evening some of the lab members went to Café Ariman in Lund and later to see the movie "The Apprentice" at #Kino in #Lund
https://www.sciencealert.com/just-one-molecule-allows-us-to-see-millions-more-colors-than-our-pets #color #vision #red #opsin #thyroid #RetinoicAcid
Our favourite #Platynereis #opsin now in the mammalian brain as a new #optogenetic tool!
"Here we ... found that the Platynereis dumerilii ciliary opsin ... is an efficient, versatile, light-activated bistable GPCR that can suppress synaptic transmission in mammalian neurons with high temporal precision in-vivo "
5/thread
Hot off the press! Move over deep homology, what about ‘deep diversity’? We review the vast variation of visual system development and physiology.
Deep Diversity: Extensive Variation in the Components of Complex Visual Systems across Animals
Understanding the molecular underpinnings of the evolution of complex (multi-part) systems is a fundamental topic in biology. One unanswered question is to what the extent do similar or different genes and regulatory interactions underlie similar complex systems across species? Animal eyes and phototransduction (light detection) are outstanding systems to investigate this question because some of the genetics underlying these traits are well characterized in model organisms. However, comparative studies using non-model organisms are also necessary to understand the diversity and evolution of these traits. Here, we compare the characteristics of photoreceptor cells, opsins, and phototransduction cascades in diverse taxa, with a particular focus on cnidarians. In contrast to the common theme of deep homology, whereby similar traits develop mainly using homologous genes, comparisons of visual systems, especially in non-model organisms, are beginning to highlight a “deep diversity” of underlying components, illustrating how variation can underlie similar complex systems across taxa. Although using candidate genes from model organisms across diversity was a good starting point to understand the evolution of complex systems, unbiased genome-wide comparisons and subsequent functional validation will be necessary to uncover unique genes that comprise the complex systems of non-model groups to better understand biodiversity and its evolution.