Tribute to Béla Novák (1956–2025) by Alexis Barr, Francis Barr and John Tyson, in J Cell Sci.
Béla Novák (1956–2025) – a legacy of scientific discovery and personal charm
In February 2025, we were shocked and deeply saddened to hear about the death of Béla Novák, Professor in Integrative Systems Biology at the University of Oxford, UK, and a beloved grandfather, father, husband, friend and colleague (Fig. 1). With his fantastic mind, irrepressible sense of humour, collegiality and dogged determination to get to the heart of a problem, Béla was absolutely one of a kind. Missing him greatly, we have tried to capture here some of the ‘essence of Béla’ so that everyone might know who he was and what he achieved.After studying mathematics and physics at the Kolcsey Ferenc Gimnazium in Budapest, Hungary, Béla matriculated in the biological engineering program at the Budapest University of Technology and Economics (BME) and Eötvös Loránd University (ELTE), Hungary. There he was introduced to experimental studies of cell growth and division by Dr Elemér László. After graduation, he began his academic career in the Department of Agricultural and Chemical Technology at BME, where he specialized in microbial engineering. Crucial to his career development was a research leave he took in the 1980s at the University of Edinburgh, UK, to study yeast physiology in Murdoch Mitchison's lab, which culminated in a joint paper on periodic bursts of CO2 production in fission yeast cells that persist after the cell division cycle is blocked (Novak and Mitchison, 1986). Béla went on to publish six more articles with Murdoch on periodic CO2 production and oxygen consumption in fission yeast.In 1991, Béla's career took another turn when he began collaborating with John Tyson (Virginia Tech, USA) on mathematical modelling of cell cycle control mechanisms (Fig. 2). Béla turned from laboratory experimentation to mathematical modelling in part because he did not have the resources in Hungary to compete with the molecular-genetics revolution sweeping through western Europe at that time, but more so because he was sure he could make a unique and lasting impression on the field by using mathematical models to explore mechanistic hypotheses in a disciplined and productive way. As he explained in an interview with István Hargittai, “I began my career as an experimental biologist, but when the molecular details of cells came to light, it was obvious to me that I should apply the mathematical and kinetic knowledge and methods that I had taught myself as a student. This was something completely new at the time” (Hargittai István, 2007). Béla's first paper together with John (Novak and Tyson, 1993), on a mathematical model of M-phase control in frog egg extracts and embryos, is now considered a classic in molecular systems biology.Over the next 14 years, Béla and John co-advised a growing team of graduate students and postdocs who cycled between Budapest and Blacksburg, Virginia, USA. Most of their students have gone on to successful careers in science and academia in Hungary, in wider Europe and in the USA. As attested by his students, Béla was a passionate teacher and mentor. Working closely with his students on their projects, he taught them to focus on the biology of the problem and showed them how to build, analyse and simulate realistic and useful mathematical models. He was a kind and encouraging mentor, but, at the same time, always demanding scientific excellence and integrity.In 2007, Béla moved to the University of Oxford's Department of Biochemistry as the Professor of Integrative Systems Biology. Béla's great passion in Oxford, as well as in Budapest, was to understand the molecular mechanisms controlling the DNA-division cycle in eukaryotic cells. He was convinced that mathematical modelling and computer simulations would play an irreplaceable role in understanding experimental studies of this control system by biochemists, geneticists and physiologists. As a theoretician, he ventured across eukaryotic kingdoms, modelling cell cycle regulation in frog and fly embryos, fission and budding yeast cells, green algae and mammalian cells. Although there were, at first, many nay-sayers of his mathematical modelling approach, it is fair to say that Béla won over many of the sceptics with his insight, expertise and charm.From 2010–2015, Béla was a key figure in the EU-funded MitoSys consortium (http://www.mitosys.org/), which took on the numerous challenges of understanding mitosis from a systems biology perspective. In 2014, he led a successful bid for a UK Biotechnology and Biological Sciences Research Council (BBSRC) sLoLa award to understand cell cycle transitions in mammalian cells by combining quantitative single-cell imaging and modelling. The award brought together labs working on G1/S (Chris Bakal), G2/M (Helfrid Hochegger), the spindle assembly checkpoint (Ulrike Gruneberg) and mitotic exit (Francis Barr), all centred on the modelling expertise of Béla and his team. This fantastic team effort was immensely productive over its 5-year term, delivering dozens of highly cited papers (Barr et al., 2016, 2017; Cundell et al., 2013, 2016; Hayward et al., 2019; Heldt et al., 2018; Hutter et al., 2017; Rata et al., 2018) and providing a wonderful training opportunity for the students and postdocs involved. The constructive feedback between theoreticians and experimentalists in this project confirmed Béla's vision of a new paradigm in molecular cell biology with mathematical models playing a central role (Tyson and Novak, 2020).Béla worked on almost all aspects of cell cycle control over his career. His work with John Tyson and their joint students focussed on frog eggs (Novak and Tyson, 1993), fission yeast (Novak and Tyson, 1995, 1997; Novak et al., 2001), budding yeast (Chen et al., 2000, 2004; Ciliberto et al., 2003), mammalian cells (Novák and Tyson, 2004) and fly embryos (Calzone et al., 2007). Later work focused on general principles of cell cycle control (Hopkins et al., 2017; Kapuy et al., 2009; Novak and Tyson, 2022; Novak et al., 2007; Tyson and Novak, 2008; Verdugo et al., 2013) and on stochastic modelling of cell cycle variability (Barik et al., 2010). In addition to the collaborations mentioned previously, Béla also had productive collaborations with Frank Uhlmann on budding yeast cell division (López-Avilés et al., 2009; Queralt et al., 2006; Vinod et al., 2011), with Tim Hunt on regulated phosphatases in mitotic control (Domingo-Sananes et al., 2011), with Damien Coudreuse on a ‘minimal cell cycle network’ (Gérard et al., 2015), with Racquel Oliveira on the spindle assembly checkpoint (He et al., 2011; Mirkovic et al., 2015; Zhang et al., 2013), with Satoru Mochida on phosphatase regulation (Mochida et al., 2016) and with Wolfgang Zachariae on meiotic prophase (Okaz et al., 2012).In May, colleagues, collaborators, friends and family gathered in Oxford to celebrate Béla's life, in what should have been his retirement symposium. Béla's irrepressible spirit was hugely missed, but everyone did their best to capture Béla's character; the day was filled with laughter as people shared their memories of meeting and working with Béla. The day was fittingly opened by John Tyson, who captured their scientific and personal journeys over 30-plus years of working together. John was followed by talks from Paul Nurse, Frank Uhlmann, Tim Hunt, Alexis Barr, Helfrid Hochegger, Kim Nasmyth, Damien Coudreuse, Ulrike Gruneberg, Raquel Oliviera, Wolfgang Zachariae and Francis Barr. The talks reflected the reach of Béla's infectious enthusiasm for modelling cell cycle control, and every talk celebrated how much fun it was to work with him. There was much discussion of how determined and hard-working Béla was. Many of us received emails at 3 am after Béla had made a breakthrough on a problem that had been burning away at him. Meetings with Béla could last hours as problems were explored from all angles and whiteboards were filled with influence diagrams and equations. He had the amazing ability to recall minute details from papers published decades ago and redraw data that had been shown in ‘Fig. 3C’ from said paper.Béla was an unforgettable character. Everyone who met him was immediately charmed by his warm and jovial nature. He treated everyone with equal respect and kindness, whether you were a student or postdoc, a secretary or security staff, a junior lecturer or senior professor, a Fields Medallist or Nobel Laureate. Béla was genuinely interested in everyone, and he took the time to show it. He was also an inveterate teaser! But it was always gentle, good-natured teasing, meant to put you at ease and to show that he recognized and valued you. Although Béla did not shrink from correcting anyone, there was always a twinkle in his eye so you wouldn't feel too chagrined. Béla had this rare skill – to challenge people while making them feel comfortable – that undoubtedly made him a great collaborator to many.Béla Novák's scientific legacy will live on for years to come (Fig. 3). But even more important to us is his personal legacy of warmth, kindness, humour, integrity, encouragement and basic humanity. Remembrances“A vibrant, generous, amusing man, one I will always remember, memories I will always cherish.” Paul Nurse“Learning from Béla not to take existing dogma for granted has ever since shaped my path.” Frank Uhlmann“With his mathematical expertise, his intimate knowledge of the literature, and his direct experience of cell cycle experimentation in Murdoch's lab, Béla [was] ideally placed to start formulating what fast became the grammar of cell cycle dynamics.” Kim Nasmyth“He was not only an exceptional researcher but also a truly kind and generous colleague who cared deeply for all of us.” Daniel Gerlich“His influence shaped my scientific path and thinking in ways I'm still discovering.” Raquel Oliveira“I learned [from Béla] how to build an ODE model, how to drink palinka the proper way, and how to lose a table tennis game with panache.” Laurence Calzone“…there is no one who conveyed such a contagious enthusiasm for everything!” Damien Coudreuse“Reviewing data with Béla was often the highpoint of my week, and talking to Béla always challenged and expanded my thinking, but most of all, was always fun.” Ulrike GrunebergBéla embodied “an endless curiosity about biology and math, mixed with an unapologetic openness in the pursuit of his passions, and a life lived for his family, friends and colleagues.” Stefan HeldtBéla's diagrams “carried a deeply satisfying message – that it was great to be a scientist studying cell division, that one had colleagues who felt the same, that one had understood a bit about the living world and its beautiful principles.” Wolfgang Zachariae“He had this uncanny ability to balance warmth with honesty, to challenge me without ever making me doubt myself.” Madhu Srinivasan“His first grandchild, Lily, was born during our sLoLa collaboration, and – I seem to recall – he was so excited that he even tried to change the name of the grant from LoLa to LiLy…” Alexis Barr“Béla's important contributions to science will endure, but he will be remembered just as much for his kindness, integrity, and the profound influence he had on those around him.” Francis Barr“I remember waiting at the airport to pick him up, wondering what this guy would be like. I expected a short, dapper, well-groomed Hungarian in a dark Russian suit, but the guy who came up to me was about my height, in jeans and a scruffy sweater, with a mop of unruly hair and a big smile. ‘This is gonna be interesting!’ I remember thinking. Little did I know, at the time!” John Tyson
Paracetamol disrupts early embryogenesis by cell cycle inhibition
https://academic.oup.com/humrep/advance-article/doi/10.1093/humrep/deaf116/8234396
#HackerNews #Paracetamol #Embryogenesis #CellCycle #Inhibition #Research #HealthScience
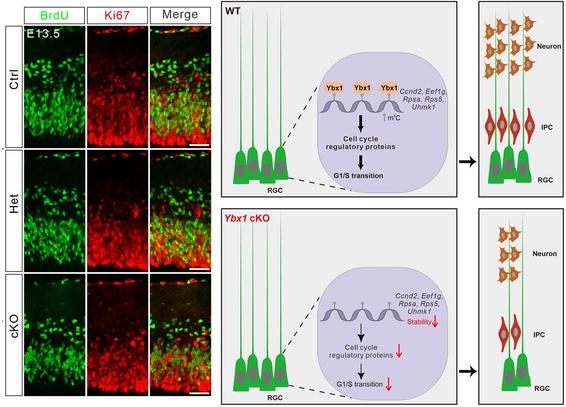
Interferon regulatory factor 4 mediates nonenzymatic IRE1 dependency in multiple myeloma cells
Multiple myeloma often requires IRE1 to sustain malignant growth, but the mechanisms remain unclear. Here, the authors show that IRE1 promotes cell-cycle progression via the transcription factor IRF4, paving the path for new therapeutic strategies.
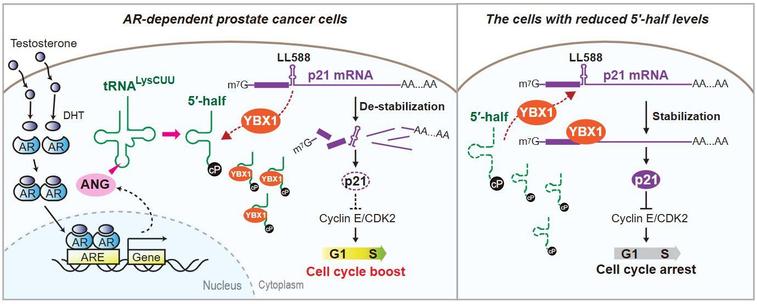
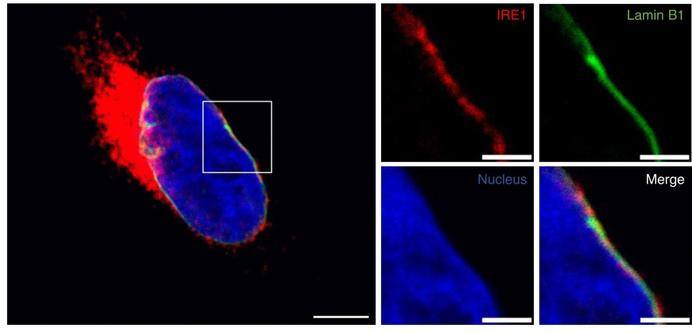
A nonenzymatic dependency on inositol-requiring enzyme 1 controls cancer cell cycle progression and tumor growth
ER-resident IRE1 is known to play a role in cancer development through its enzymatic activation of XPB1 transcription factors and of regulated RNA decay. This study shows that some cancer cells also have an unexpected nonenzymatic dependency on IRE1 to promote cell cycle progression and tumor growth.
FLCCR is a fluorescent reporter system that quantifies the duration of different cell cycle phases at the single-cell level in fission yeast
Current methods to study cell cycle progression in yeast overlook variations in cell size across different strains. This study develops an imaging reporter system called FLCCR that can quantify the duration of each cell cycle phase in single cells and allows the separation of cell size as an independent variable from cell cycle progression.
From post 5,
Amino acids that are enriched/depleted next to the phospho-site reflect which protein kinase enzymes did the phosphorylation.
Phospho-dawn sees acid-directed sites being phosphorylated (D and E enriched at ZT0),
but by evening it's #CellCycle kinases guided by Proline (P at ZT12-16). 😎
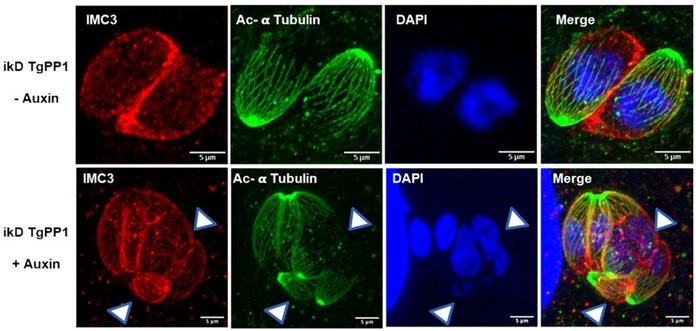
PP1 phosphatase controls both daughter cell formation and amylopectin levels in Toxoplasma gondii
Rapid division and biomass production are important for apicomplexan parasite virulence. This study shows that the phosphatase TgPP1 in Toxoplasma gondii regulates the cell cycle and amylopectin accumulation by dephosphorylating inner membrane complex and starch-binding domain proteins.