https://www.biorxiv.org/content/10.64898/2026.04.29.721761v1?rss=1
#Drosophila #Sleep
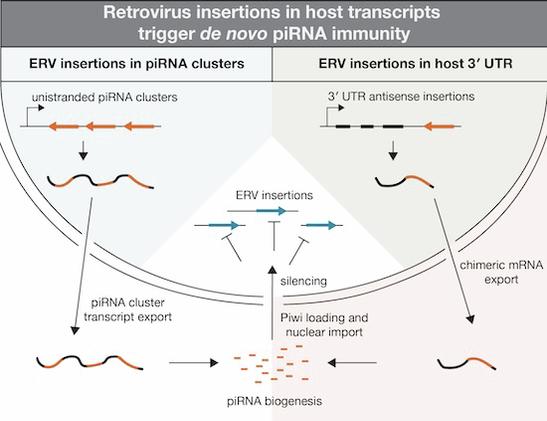
How host organisms adapt their defense systems to newly invading transposable elements remains poorly understood. Here, we show how Drosophila melanogaster acquired PIWI-interacting RNA (piRNA)-mediated immunity against the endogenous retrovirus Tirant. We uncover two distinct modes of de novo piRNA biogenesis by combining genetics, small RNA profiling, and population genomics. The primary route involves antisense insertions into the flamenco cluster, a master locus for transposon control. Unexpectedly, a second, equally potent mechanism arises from antisense Tirant insertions within host gene 3′ UTRs. This process requires host gene transcription but is independent of host gene identity. Our findings challenge prevailing models that tie piRNA precursor specification to genomic origin or nuclear RNA processing context. Instead, they reveal a flexible mechanism that turns a critical vulnerability of transposons into an advantage for the host. When transposition occurs into host gene exons, chimeric antisense transcripts are exported to the cytoplasm, inadvertently initiating piRNA production and enabling rapid, adaptive genome defense against new invaders.
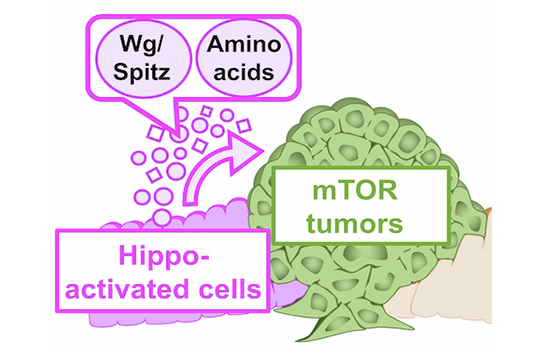
The Hippo pathway is a tumor suppressor pathway, and most related studies have indicated that its inhibition leads to tumorigenesis. However, recent studies have suggested that the activated Hippo pathway can promote tumorigenesis in certain contexts. Here, we demonstrate that the activated Hippo pathway induces non-cell-autonomous tumorigenesis, characterized by tumor markers in the Drosophila wing epithelium. This suggests that Hippo-activated cells behave similarly to “oncogenic niche cells.” We find that Hippo-activated cells induce Dronc-Wingless/Spitz signaling in the hinge/ventral notum region, which causes tumorigenesis. Moreover, we identify the amino acid transporters Sat1/2, which are implicated in amino acid incorporation and function redundantly with the growth factors Wingless and Spitz to facilitate non-cell-autonomous tumorigenesis.