Postdoc in Environmental Regulation of Reproduction via RNA Modifications
Post a job in 3min, or find thousands of job offers like this one at jobRxiv!

Postdoc in Environmental Regulation of Reproduction via RNA Modifications
Post a job in 3min, or find thousands of job offers like this one at jobRxiv!
What is the role of m5C
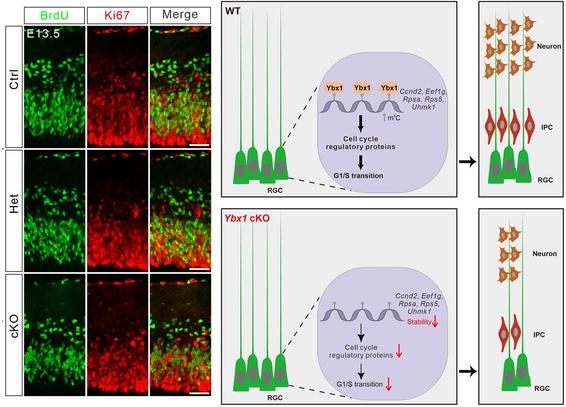
#RNAmodifications during neural development? This study shows that m5C reader protein Ybx1 plays a crucial role in promoting
#CellCycle progression of embryonic cortical
#ProgenitorCells, essential for proper cortical development
@PLOSBiology https://plos.io/3SUN4we📚 Our latest review article "Beyond the Anticodon: tRNA Core Modifications and Their Impact on Structure, Translation, and Stress Adaptation" is out!
🎯 Read more about tRNA core modifications, which should not be overlooked in favor of their anticodon-loop counterparts. #tRNA #RNAmodifications #Translation #StressAdaptation
https://www.mdpi.com/2073-4425/15/3/374

Beyond the Anticodon: tRNA Core Modifications and Their Impact on Structure, Translation and Stress Adaptation
Transfer RNAs (tRNAs) are heavily decorated with post-transcriptional chemical modifications. Approximately 100 different modifications have been identified in tRNAs, and each tRNA typically contains 5–15 modifications that are incorporated at specific sites along the tRNA sequence. These modifications may be classified into two groups according to their position in the three-dimensional tRNA structure, i.e., modifications in the tRNA core and modifications in the anticodon-loop (ACL) region. Since many modified nucleotides in the tRNA core are involved in the formation of tertiary interactions implicated in tRNA folding, these modifications are key to tRNA stability and resistance to RNA decay pathways. In comparison to the extensively studied ACL modifications, tRNA core modifications have generally received less attention, although they have been shown to play important roles beyond tRNA stability. Here, we review and place in perspective selected data on tRNA core modifications. We present their impact on tRNA structure and stability and report how these changes manifest themselves at the functional level in translation, fitness and stress adaptation.

Imaging mRNA modifications in situ
There are at least 170 known chemical modifications on RNA molecules in all organisms. Most of these modifications are on tRNAs, rRNAs and small RNAs. But over the past two decades, researchers fou…