Our preprint is up! Take a look if you like kinesin, microtubule transport, single molecule FRET or biophysics. It’s a bit of shift to how we’ve been used to thinking about kinesin-1 regulation as we try to get a handle on its conformational dynamics. Super proud of to have this milestone out in the world and all the hard work it represents from everyone involved.
#kinesin #microtubules #biophys #smFRET #SingleMolecule #AxonalTransport https://biologists.social/@TwelvetreesLab/113691748206473977Kinesin-1 is highly flexible and adopts an open conformation in the absence of cargo
http://biorxiv.org/cgi/content/short/2024.12.20.629623v1?rss=1
Does anyone have strong opinions on what the best/most sensitive low volume spectrophotometer is? Looking to quantify low concentrations of fluorescently labelled protein, not just DNA etc
#biochem #biophys #singlemolecule
Updated our #singlemolecule preprint on "Enhancing the Applied Force and Range of Axial Optical Tweezers”
https://www.biorxiv.org/content/10.1101/2023.04.21.537890v2

Science Jobs - Find science and research jobs - 3,034 jobs
The international job board for scientists, by scientists. Find science jobs in academia or industry: MSc, PhD, Postdoc, Scientist, Faculty and more!
Depixus wins SLAS2024 Ignite Award for MAGNA™ technology - Depixus
12th February 2024 Paris, France - Depixus CEO Gordon Hamilton Receives The SLAS2024 Ignite Award From Judge Severine Tamas-Lhoustau, CEO Of Novoptim.
Our work has made the inside cover! Check out "
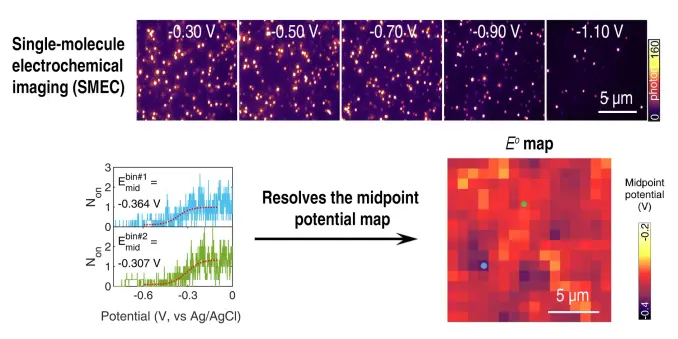
#singlemolecule #electrochemical imaging resolves the midpoint potentials of individual fluorophores on nanoporous antimony-doped tin oxide" in the latest issue of
@chemicalscience!
https://doi.org/10.1039/d3sc05293a @washuengineers #WashUESE #WUSTLCongratulations to Prof. Jin Lu, now at the National Center for Nanoscience, whose work on #singlemolecule #electrochemical imaging is now online in Chemical Science. Happy New Year!
#SMLM #WashU #WUSTL @chemicalscience #ChemicalScience
"Single-Molecule Electrochemical Imaging Resolves the Midpoint Potentials of Individual Fluorophores on Nanoporous Antimony-Doped Tin Oxide"
Chemical Science, 2024
https://doi.org/10.1039/D3SC05293A
If you are a #microscopy maker, passionate about our research, and envision ending your days with such views, contact us! We're opening a new #postdoc position.
#SPADarray #singlephotonmicroscopy #singlemolecule #lightsheetmicroscopy #BrigthEyes_ERC

Episode Gallery | The Master Minds Podcast
new citation:
`Using salt tolerance compartmentalized self-replication (stCSR) and a #microfluidic platform, we obtained 11 mutant sites with enhanced salt tolerance attributes. #Sequencing and #biochemical analyses revealed that the substitution of conserved amino acids such as G197D, Y369E, T372N, and I378R plays a critical role in maintaining the #processivity of exonuclease-deficient #phi29 #polymerase under high salt conditions.`
https://www.frontiersin.org/articles/10.3389/fmicb.2023.1267196/full
#nanopore #DNA #singleMolecule

Unraveling the salt tolerance of Phi29 DNA polymerase using compartmentalized self-replication and microfluidics platform
In Phi29-α–hemolysin (α-HL) nanopore sequencing systems, a strong electrochemical signal is dependent on a high concentration of salt. However, high salt concentrations adversely affect polymerase activity. Sequencing by synthesis (SBS) requires the use of phi29 polymerase without exonuclease activity to prevent the degradation of modified nucleotide tags; however, the lack of exonuclease activity also affects polymerase processivity. This study aimed to optimize phi29 polymerase for improved salt tolerance and processivity while maintaining its lack of exonuclease activity to meet the requirements of nanopore sequencing. Using salt tolerance compartmentalized self-replication (stCSR) and a microfluidic platform, we obtained 11 mutant sites with enhanced salt tolerance attributes. Sequencing and biochemical analyses revealed that the substitution of conserved amino acids such as G197D, Y369E, T372N, and I378R plays a critical role in maintaining the processivity of exonuclease-deficient phi29 polymerase under high salt conditions. Furthermore, Y369E and T372N have been identified as important determinants of DNA polymerase binding affinity. This study provides insights into optimizing polymerase processability under high-salt conditions for real-time polymerase nanopore sequencing, paving the way for improved performance and applications in nanopore sequencing technologies.