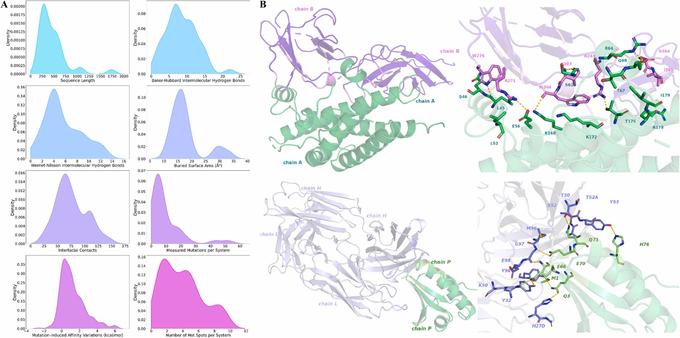
🚀 What if we could precisely target the CAPON–nNOS axis — a pathway long considered undruggable?
🔗 Targeting CAPON to modulate the CAPON–NOS Axis: a computational approach. Computational and Structural Biotechnology Journal, DOI: https://doi.org/10.1016/j.csbj.2025.11.001
📚 CSBJ: https://www.csbj.org/
#DrugDiscovery #ComputationalBiology #AlzheimersResearch #Neuroscience #ProteinProteinInteractions #VirtualScreening #MolecularDynamics #AIinDrugDiscovery
💡 AI can predict protein structures — but can it predict how they behave?
🔗 AlphaFold3 prediction of protein-protein complex: Is it ready for thermodynamic analysis?. Computational and Structural Biotechnology Journal, DOI: https://doi.org/10.1016/j.csbj.2025.10.015
📚 CSBJ: https://www.csbj.org/
#AlphaFold3 #ProteinDesign #MolecularModeling #ComputationalBiology #StructuralBiology #Thermodynamics #ProteinProteinInteractions #Bioinformatics
Distinct Functions of the PH Domain in BCR/ABL p210 Isoform: Interaction with Cytoskeletal and Membrane Remodeling Proteins -
#BCRABL #cortactin #FBP17 #PHdomain #CML #GSTpulldown #proteinproteininteractions -
https://link.springer.com/article/10.3103/S0095452725020045
Distinct Functions of the PH Domain in BCR/ABL p210 Isoform: Interaction with Cytoskeletal and Membrane Remodeling Proteins - Cytology and Genetics
Abstract The BCR/ABL fusion protein, generated by the Philadelphia chromosome translocation, drives chronic myelogenous leukemia (CML) and other myeloproliferative disorders. The p210 isoform includes a pleckstrin homology (PH) domain absent in the p190 isoform, which is linked to acute lymphoblastic leukemia (ALL). This structural difference may underlie the distinct subcellular localization and signaling profiles of the two isoforms. Here, we investigate the role of the PH domain of BCR in interactions with cortactin and FBP17, proteins involved in cytoskeletal remodeling and membrane dynamics. Using GST-pulldown assays and western blotting, we demonstrated direct interactions between the PH domain of BCR and both cortactin and FBP17. Colocalization studies, supported by confocal and STED microscopy, revealed that cortactin colocalizes with the PH domain of BCR in the centrosomal and perimembrane regions of cells. Notably, the SH3 domain of cortactin was not required for this interaction, but full-length cortactin was essential, suggesting that other domains mediate binding. These findings highlight the role of the PH domain in directing BCR/ABL to the centrosome, where it interacts with cortactin to potentially influence actin dynamics and vesicular trafficking. This centrosomal localization may spatially restrict the constitutive tyrosine kinase activity of ABL, contributing to the less aggressive phenotype of p210-associated CML compared to p190-driven ALL. Understanding the role of the PH domain as a key structural difference between p210 and p190 is critical for elucidating the molecular basis of BCR/ABL-mediated leukemogenesis. Future studies will explore the phosphorylation of cortactin and FBP17 by ABL kinase and the domains responsible for these interactions.

Janus-Like Nature Leads to Unexpected Protein Oligomer Formation - ChemistryViews
Weak self-interactions lead to the formation of small but stable oligomers

AlphaProteo generates novel proteins for biology and health research
New AI system designs proteins that successfully bind to target molecules, with potential for advancing drug design, disease understanding and more.
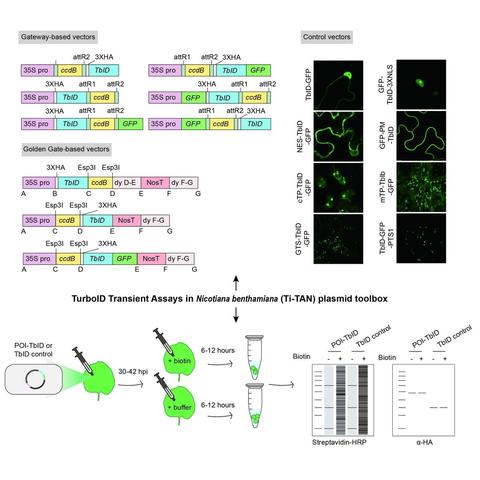
Huang Tan et al. describe their new Ti-TAN
#plasmid toolbox for TurboID-based proximity labeling assays in N. benthamiana. Read the full paper here!
#PlantScience #OpenAccess #proteinproteininteractions #TurboID https://onlinelibrary.wiley.com/doi/10.1111/jipb.13610A new proteomic workflow establishing interdependencies between protein
#phosphorylation & complex formation is applied to the signal integrator & Hippo pathway effector YAP1 ➡️
http://bit.ly/3mJ4W0t Matthias Gstaiger
@eth_en #proteomics #interactome #ProteinProteinInteractions #CellSignalingNews &Views by
P Kastritis on the recent study from the Rappsilber lab identifying bacterial PPIs in their cellular context by combining whole-cell crosslinking, co-fractionation
#massspectrometry & AI-based PPI structure prediction ➡️
http://bit.ly/3FfcIVY#proteomics #ProteinProteinInteractions #SystemsBiology #AlphafoldAn integrative approach using crosslinking
#massspectrometry (MS), co-fractionation MS &
#Alphafold-Multimer discovers new protein complexes & their topologies in B.subtillis ➡️
http://bit.ly/3kpjocUfrom Juri Rappsilber (TU Berlin), Jörg Stülke (Uni Goettingen)
#proteomics #SystemsBiology #ProteinProteinInteractionsABOLISH: an optimised biotin-ligation method for discovering stable & transient
#ProteinProteinInteractions in yeast ➡️
https://www.embopress.org/doi/full/10.15252/msb.202211084 from Maya Schuldiner's lab, Weizmann Institute
#SystemsBiology #Scerevisiae