💧 Is your MD simulation lying to you because of the water model you picked?
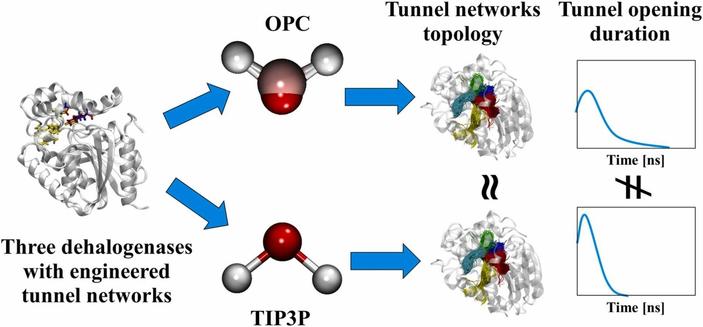
🔗 Impact of water models on the structure and dynamics of enzyme tunnels. Computational and Structural Biotechnology Journal, DOI: https://doi.org/10.1016/j.csbj.2024.10.051
📚 CSBJ: https://www.csbj.org/
#ComputationalBiology #MolecularDynamics #Enzymology #StructuralBiology #ProteinEngineering #Biophysics #WaterModels #EnzymeTunnels #MolecularSimulation #ProteinDynamics