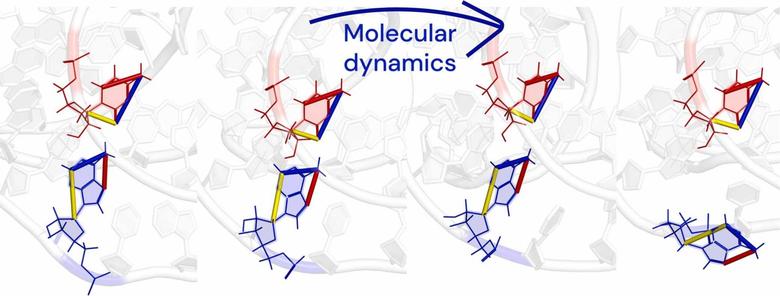
🧬 When does molecular dynamics improve RNA models? Insights from CASP15 and practical guidelines. Computational and Structural Biotechnology Journal, DOI: https://doi.org/10.1016/j.csbj.2025.10.003
📚 CSBJ: https://www.csbj.org/
#RNA #StructuralBiology #CASP15 #ComputationalBiology #Bioinformatics #MolecularSimulation #MolecularDynamics #Biophysics #MolecularModeling