Another thought-provoking
#MedChem #SciComm story by Stan Van Boeckel.
On the example of Sitagliptin, he explains the power of minimizing molecules and looking for the highest ligand efficiency in
#DrugDiscovery.
www.linkedin.com/posts/stan-v...
#ChemSky #ChemBio #SmallMolecules #Science #MedicinesPart 29 — Sometimes “smaller i...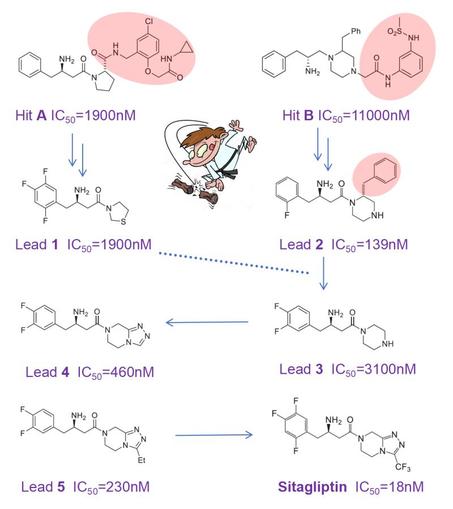
Part 29 — Sometimes “smaller is elegant” in drug discovery.
In early discovery, medicinal chemists often chase potency by making molecules larger and more lipophilic. Yet potency should always be… | Stan Van Boeckel
Part 29 — Sometimes “smaller is elegant” in drug discovery.
In early discovery, medicinal chemists often chase potency by making molecules larger and more lipophilic. Yet potency should always be interpreted in the context of molecular weight and lipophilicity. Ligand‑efficiency metrics normalize potency to size and logP, helping chemists distinguish truly efficient hits and leads from bulky or greasy molecules that only appear potent but usually bring ADMET liabilities. The real challenge is to think broadly, so consider to make your hit or lead smaller during optimization, or 'just do it'.
The discovery of sitagliptin illustrates this beautifully: a blockbuster drug that is smaller than the original screening hits. Sitagliptin (approved 2006) is the first DPP‑4 inhibitor that slows the rapid degradation of GLP‑1 and GIP, thereby lowering plasma glucose. In Merck’s HTS campaign, compounds A and B emerged as low‑potency hits. Here comes the twist: instead of growing these hits, the team chopped off large moieties and explored whether the resulting fragments could be optimized with small substituents such as fluorines. Surprisingly, this yielded compact leads 1 and 2 with improved potency, while selectivity across DPP family members and oral bioavailability in rat were continuously monitored.
Lead 2 was then chopped again-taking inspiration from lead 1, which lacked its extra phenyl ring-to give weak lead 3. Accepting a 20‑fold drop in IC₅₀ seems counterintuitive, but it opened the door to expand the piperazine and create a new heterocycle. This delivered more potent lead 4, followed by lead 5. Further fluorination of 5 improved oral exposure and ultimately delivered the blockbuster sitagliptin.
This small, well‑balanced molecule (MW 407; IC₅₀ 18 nM; oral bioavailability 87%; t½ 12 h; rapid absorption; logP 1.5; high selectivity; medium solubility; 5 rotatable bonds; HA/HD 4/1; non‑toxic) is a superb example of optimizing property space without molecular obesity. A remarkable achievement by Malcolm MacCoss, Ph.D., FRSC and his team at Merck.

Part 29 — Sometimes “smaller is elegant” in drug discovery.
In early discovery, medicinal chemists often chase potency by making molecules larger and more lipophilic. Yet potency should always be… | Stan Van Boeckel
Part 29 — Sometimes “smaller is elegant” in drug discovery.
In early discovery, medicinal chemists often chase potency by making molecules larger and more lipophilic. Yet potency should always be interpreted in the context of molecular weight and lipophilicity. Ligand‑efficiency metrics normalize potency to size and logP, helping chemists distinguish truly efficient hits and leads from bulky or greasy molecules that only appear potent but usually bring ADMET liabilities. The real challenge is to think broadly, so consider to make your hit or lead smaller during optimization, or 'just do it'.
The discovery of sitagliptin illustrates this beautifully: a blockbuster drug that is smaller than the original screening hits. Sitagliptin (approved 2006) is the first DPP‑4 inhibitor that slows the rapid degradation of GLP‑1 and GIP, thereby lowering plasma glucose. In Merck’s HTS campaign, compounds A and B emerged as low‑potency hits. Here comes the twist: instead of growing these hits, the team chopped off large moieties and explored whether the resulting fragments could be optimized with small substituents such as fluorines. Surprisingly, this yielded compact leads 1 and 2 with improved potency, while selectivity across DPP family members and oral bioavailability in rat were continuously monitored.
Lead 2 was then chopped again-taking inspiration from lead 1, which lacked its extra phenyl ring-to give weak lead 3. Accepting a 20‑fold drop in IC₅₀ seems counterintuitive, but it opened the door to expand the piperazine and create a new heterocycle. This delivered more potent lead 4, followed by lead 5. Further fluorination of 5 improved oral exposure and ultimately delivered the blockbuster sitagliptin.
This small, well‑balanced molecule (MW 407; IC₅₀ 18 nM; oral bioavailability 87%; t½ 12 h; rapid absorption; logP 1.5; high selectivity; medium solubility; 5 rotatable bonds; HA/HD 4/1; non‑toxic) is a superb example of optimizing property space without molecular obesity. A remarkable achievement by Malcolm MacCoss, Ph.D., FRSC and his team at Merck.
Isomorphic Labs Prepares AI-Designed Drugs for Initial Human Trials
📰 Original title: AI-Designed Drugs by a DeepMind Spinoff Are Headed to Human Trials
🤖 IA: It's not clickbait ✅
👥 Usuarios: It's not clickbait ✅
View full AI summary: https://killbait.com/en/isomorphic-labs-prepares-ai-designed-drugs-for-initial-human-trials/?redirpost=c1ae24f9-16bf-4d11-9945-74a5c43d86bb
#health #ai #drugdiscovery #biotech

Isomorphic Labs Prepares AI-Designed Drugs for Initial Human Trials
Isomorphic Labs, a biotech spinoff from Google DeepMind, is preparing to enter human trials with drugs designed using its AI technology, AlphaFold. Announced at WIRED Health in London…
Isomorphic Labs Prepares AI-Designed Drugs for Initial Human Trials
📰 Original title: AI-Designed Drugs by a DeepMind Spinoff Are Headed to Human Trials
🤖 IA: It's not clickbait ✅
👥 Usuarios: It's not clickbait ✅
View full AI summary: https://killbait.com/en/isomorphic-labs-prepares-ai-designed-drugs-for-initial-human-trials/?redirpost=c1ae24f9-16bf-4d11-9945-74a5c43d86bb
#health #ai #drugdiscovery #biotech

Isomorphic Labs Prepares AI-Designed Drugs for Initial Human Trials
Isomorphic Labs, a biotech spinoff from Google DeepMind, is preparing to enter human trials with drugs designed using its AI technology, AlphaFold. Announced at WIRED Health in London…
Part 27, drug discovery on an “Undruggable” Target.
K‑Ras is a small intracellular GTPase that functions as a molecular switch, activating mitogenic signaling pathways in response to growth… | Stan Van Boeckel
Part 27, drug discovery on an “Undruggable” Target.
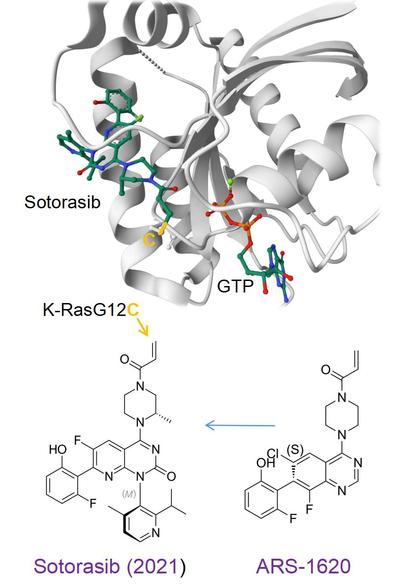
K‑Ras is a small intracellular GTPase that functions as a molecular switch, activating mitogenic signaling pathways in response to growth factors. Direct attempts to inhibit this oncogene with competitive drugs at its GTP‑binding site failed because of its picomolar affinity for GTP and the high intracellular GTP concentration. Mutations in the K‑Ras pathway, such as K‑RasG12C, render the protein constitutively active, driving uncontrolled proliferation and e.g. contributing to ~40% of K‑Ras–driven lung cancers. Until 2013, K‑Ras was considered non‑druggable, but this view shifted when the Shokat lab revealed a hidden allosteric regulatory pocket in K‑RasG12C that becomes accessible when small electrophilic molecules covalently bind the mutant cysteine. This covalent engagement displaces key “protein switches,” biases the protein toward GDP over GTP conformation and prevents Raf binding, thereby shutting down MAPK signaling.
Following this breakthrough, Amgen optimized the Shokat group’s early acrylamide fragments into an in‑vivo‑suitable tool compound, ARS‑1620. Extensive crystallography and docking guided the medicinal chemistry cycles that ultimately produced the highly decorated drug sotorasib (approved in 2021). In phase 3 trials in K‑RasG12C‑mutant lung cancer, sotorasib improved progression‑free survival compared with docetaxel, although overall survival was unchanged. That outcome is somewhat disappointing, but there is hope that real‑world drug combination strategies may yet deliver meaningful gains in overall survival.

Part 27, drug discovery on an “Undruggable” Target.
K‑Ras is a small intracellular GTPase that functions as a molecular switch, activating mitogenic signaling pathways in response to growth… | Stan Van Boeckel
Part 27, drug discovery on an “Undruggable” Target.
K‑Ras is a small intracellular GTPase that functions as a molecular switch, activating mitogenic signaling pathways in response to growth factors. Direct attempts to inhibit this oncogene with competitive drugs at its GTP‑binding site failed because of its picomolar affinity for GTP and the high intracellular GTP concentration. Mutations in the K‑Ras pathway, such as K‑RasG12C, render the protein constitutively active, driving uncontrolled proliferation and e.g. contributing to ~40% of K‑Ras–driven lung cancers. Until 2013, K‑Ras was considered non‑druggable, but this view shifted when the Shokat lab revealed a hidden allosteric regulatory pocket in K‑RasG12C that becomes accessible when small electrophilic molecules covalently bind the mutant cysteine. This covalent engagement displaces key “protein switches,” biases the protein toward GDP over GTP conformation and prevents Raf binding, thereby shutting down MAPK signaling.
Following this breakthrough, Amgen optimized the Shokat group’s early acrylamide fragments into an in‑vivo‑suitable tool compound, ARS‑1620. Extensive crystallography and docking guided the medicinal chemistry cycles that ultimately produced the highly decorated drug sotorasib (approved in 2021). In phase 3 trials in K‑RasG12C‑mutant lung cancer, sotorasib improved progression‑free survival compared with docetaxel, although overall survival was unchanged. That outcome is somewhat disappointing, but there is hope that real‑world drug combination strategies may yet deliver meaningful gains in overall survival.
Very much looking forward to today's @LED3hub Lecture by Peng Chen on "Live-cell Protein Chemistry in Health and Disease".
If you are interested, make sure to come by!
https://www.universiteitleiden.nl/en/science/led3/led3-lectures
#Chemistry #ChemBio #Cells #Proteins #DrugDiscovery
Very much looking forward to today's
@[email protected] Lecture by Peng Chen on "Live-cell Protein Chemistry in Health and Disease".
If you are interested, make sure to come by!
www.universiteitleiden.nl/en/science/l...
#ChemSky #ChemBio #Cells #Proteins #DrugDiscoveryNew Algorithmic Scrutiny Targets Molecular Migrations in Drug Development
University of Oregon's new AI tool helps drug makers test new molecules faster and cheaper. It predicts how drugs will work before lab tests.
#DrugDiscovery, #AIinPharma, #UniversityOfOregon, #MolecularModeling, #HealthcareTech
https://newsletter.tf/oregon-university-ai-speeds-up-drug-making/
This new AI tool from the University of Oregon can help create new medicines much faster than before. It uses computers to predict how molecules will behave.
#DrugDiscovery, #AIinPharma, #UniversityOfOregon, #MolecularModeling, #HealthcareTech
https://newsletter.tf/oregon-university-ai-speeds-up-drug-making/

Oregon University AI speeds up new drug making
University of Oregon's new AI tool helps drug makers test new molecules faster and cheaper. It predicts how drugs will work before lab tests.






