Many people think the history of the Earth is hard to decipher. The geological record jumbles everything up. We set out to fact-check this in my ERC Starting Grant MindTheGap. Focusing on the mode of evolution, because that is something that we can only glean from long records, longer than anything accessible to observation or experimentation today.
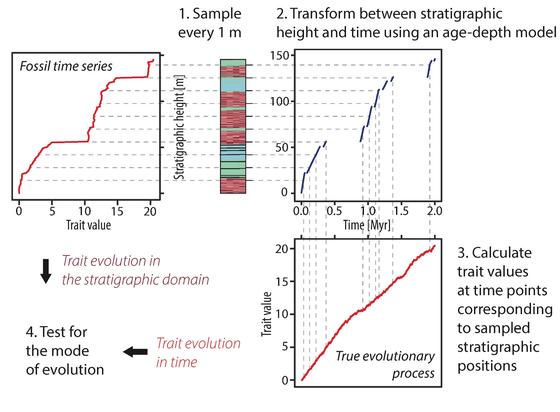
Thanks to collaboration with Peter Burgess we used forward models to simulate formation of carbonate rocks and see how completeness and stratigraphic resolution affect the reconstructions. Guess what! The geological record is 👌🏻 With a bit of understanding of #sedimentology and #physics we can test is with computer experiments and it turns out even very patchy rock strata will let us reconstruct the correct mode of evolution. This opens a lot of possibilities, especially for the #microfossil community: your data is priceless to understand #microevolution.
Check out the results at https://lnkd.in/eeiSuRwP
Also a big shoutout to an excellent review process at PCI Paleo, the thoughtful editors and reviewers, and to the lead author Niklas Hohmann for #code #reproducibility effort that meets the most stringent standards. Check out our software at https://lnkd.in/eJkwhhuh
#evolution #paleontology #paleobiology #UtrechtUniversity