Weak supervision - Strong results! 💪
Smith and team introduce Perturbational Metric Learning (PeML) to extract biological relationships from noisy high-throughput perturbational datasets.
Team effort from preLighters Benjamin Dominik Maier & Anna Foix Romero – read their preLight! 👀
#preLight 👉 https://prelights.biologists.com/highlights/similarity-metric-learning-on-perturbational-datasets-improves-functional-identification-of-perturbations/
#bioinformatics #SystemsBiology #clustering #correlations #gsea #MachineLearning
Similarity metric learning on perturbational datasets improves functional identification of perturbations - preLights
Weak supervision - Strong results! Smith and colleagues introduce Perturbational Metric Learning (PeML), a weakly supervised similarity metric learning method to extract biological relationships from noisy high-throughput perturbational datasets
Another great edition of the #GSEA in R/Bioconductor is going to an end.
Many thanks to Zuguang Gu & all attendees for this very productive week!
#GSEA #Rstats @Bioconductor
#Genomics #Bioinformatics #DataScience
I haven't heard of sparrow, by @lianos, for #gsea analysis and exploration for results, but it looks useful. In particular because of the sister #shiny package. Must explore further.
https://tomsing1.github.io/blog/posts/interactive-gene-set-results/
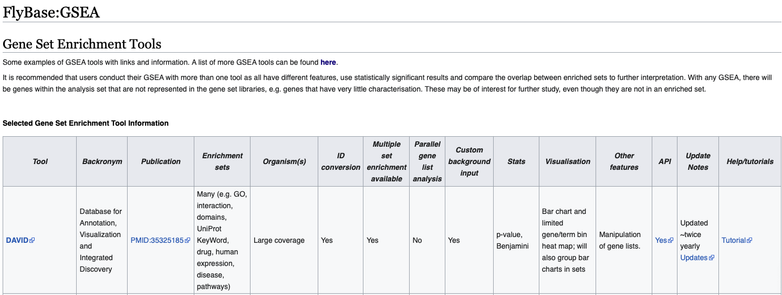
Interested in Gene Set Enrichment in R? Have a look at our new course with Dr. Zuguang Gu
If interested, please check it out: https://physalia-courses.org/courses-workshops/gse-in-r/
#Rstats #Genomics #bioconductor #DataScience #Bioinformatics #datavisualization #GSEA #CancerGenomics
https://tomsing1.github.io/blog/posts/interactive-gene-set-results/
It's a static HTML page, e.g. no server (#shiny, #dash, etc) needed. Thanks a lot to the authors of the #plotly #reactable #crosstalk and #htmlwidget tools for making this so easy #til #rstats #bioconductor #gsea #compbio #visualization @lianos