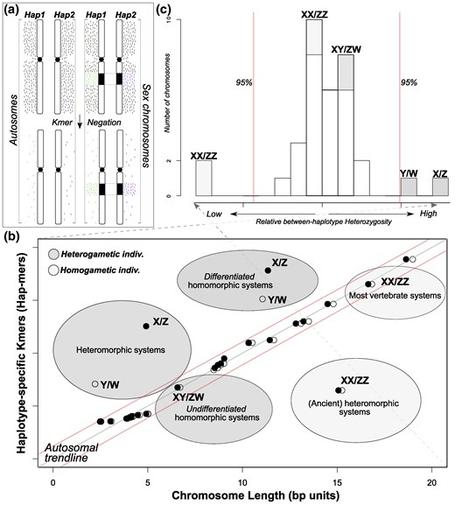
Pinto et al. present SCINKD as a framework to identify unannotated sex chromosomes and curate diploid genome assemblies from a single individual.
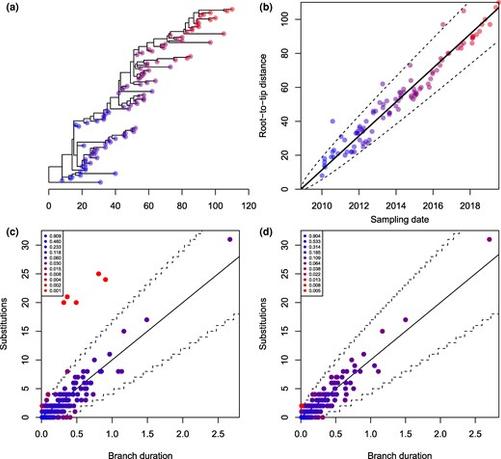
Didelot et al. present the R package DiagnoDating for diagnosing issues in a reconstructed dated phylogeny, including outlier detection, posterior predictive checking and residual analysis.
Daniel Huson introduces displacement-optimized tanglegrams (DO-tanglegrams), a new approach that applies equally to trees and rooted phylogenetic networks, performing better than cophylo on trees and then NN-tanglegram on networks.
McArthur et al. present piqtree, an easy to use, open-source Python package that provides Python script-based control of IQ-TREE’s phylogenetic inference engine.
@sishuowang & Meade introduce phyloHessian to enable the use of complex mixture substitution models in molecular dating. Empirical analysis of ancient symbiont lineages leads to a revised understanding of their host association origins.
Robbins, Liu & Kelly present RECUR, a method for identifying recurrent amino acid substitutions from multiple sequence alignments that is fast, easy to use, and scalable to thousands of sequences.
Deng et al. present TreeProfiler, a tool for automated annotation and interactive exploration of hundreds of features along large gene and species trees, with seamless summarization of mapped traits at internal nodes.
Martí-Gómez et al. developed gpmap-tools, integrating models for inference, phenotypic imputation, and error estimation from multiplex assays of variant effect data or natural sequences in the presence of genetic interactions.
Anchieri et al. benchmark the inference of selection with aDNA-like time series datasets, showing that ApproxWF can accurately estimate selection with datasets of ∼100 individuals when selection is strong.
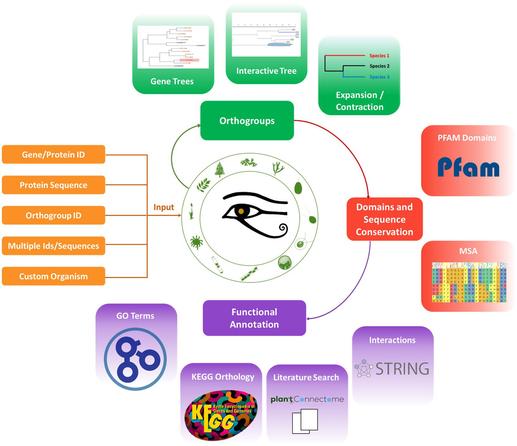
Ramos-González et al. present PharaohFUN, a web application designed for the evolutionary and functional analysis of protein sequences in photosynthetic eukaryotes, leveraging orthology relationships.