Florian Jug's lab @florianjug latest:
"MicroSplit: semantic unmixing of fluorescent microscopy data", Ashesh et al. 2026
https://www.nature.com/articles/s41592-026-03082-1
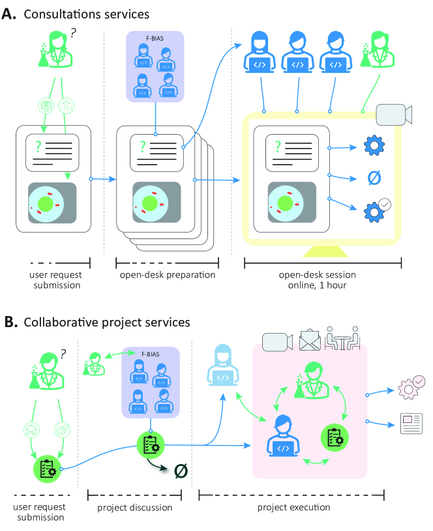
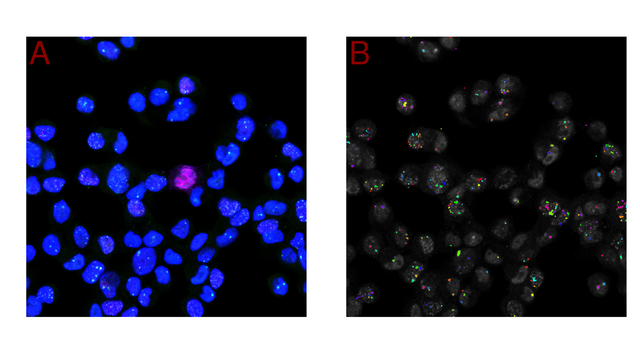
"MicroSplit separates up to four superimposed noisy structures into distinct, denoised image channels, enabling faster and more photon-efficient imaging."
"Built on Variational Splitting Encoder-Decoder networks, models a posterior distribution over solutions, allowing uncertainty-aware predictions and the estimation of spatially resolved prediction errors from posterior variability."
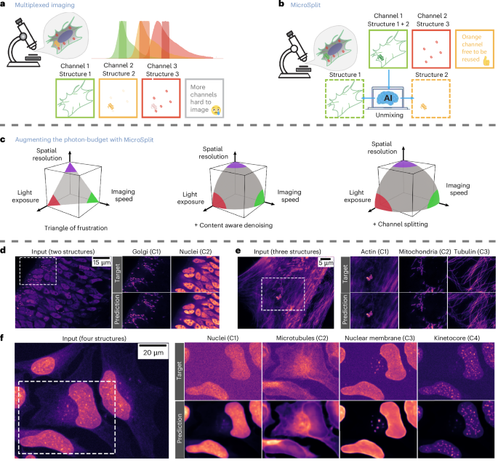
$${\bf{Micro}}{{\mathbb{S}}}{\bf{plit}}$$ Micro S plit : semantic unmixing of fluorescent microscopy data - Nature Methods
$${\rm{Micro}}{\mathbb{S}}{\rm{plit}}$$ M i c r o S p l i t is a computational unmixing method for multiplexed fluorescence imaging of up to four structures per imaging channel.