mRNA 3′UTRs chaperone intrinsically disordered regions to control protein activity
More than 2,700 human mRNA 3′UTRs have hundreds of highly conserved (HC) nucleotides, but their biological roles are unclear. Here, we show that mRNAs with HC 3′UTRs mostly encode proteins with long intrinsically disordered regions (IDRs), including MYC, UTX, and JMJD3. These proteins are only fully active when translated from mRNA templates that include their 3′UTRs, raising the possibility of functional interactions between 3′UTRs and IDRs. Rather than affecting protein abundance or localization, we find that HC 3′UTRs control transcriptional or histone demethylase activity through co-translationally determined protein oligomerization states that are kinetically stable. 3′UTR-dependent changes in protein folding require mRNA-IDR interactions, suggesting that mRNAs act as IDR chaperones. These mRNAs are multivalent, a biophysical RNA feature that enables their translation in network-like condensates, which provide favorable folding environments for proteins with long IDRs. These data indicate that the coding sequence is insufficient for the biogenesis of biologically active conformations of IDR-containing proteins and that RNA can catalyze protein folding.
### Competing Interest Statement
The authors have declared no competing interest.
Pershing Square Foundation, https://ror.org/04tce9s05
G. Harold & Leila Y. Mathers Foundation
National Institutes of Health, DP1GM123454, R35GM144046
Memorial Sloan Kettering Cancer Center, https://ror.org/02yrq0923, P30 CA008748
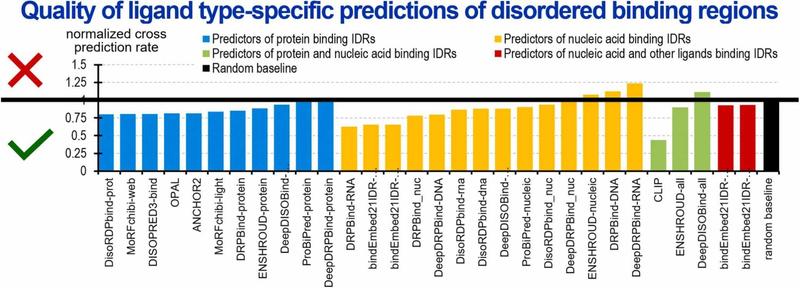
🧠 How do prediction models perform in recognizing disordered regions with specific binding functions?
🔗 Evaluation of predictions of disordered binding regions in the CAID2 experiment. Computational and Structural Biotechnology Journal, DOI: https://doi.org/10.1016/j.csbj.2024.12.009
📚 CSBJ: https://www.csbj.org/
#Bioinformatics #ProteinDisorder #CAID2 #IntrinsicallyDisorderedProteins #LigandBinding #ComputationalBiology #AIinScience #DeepLearning #MolecularBiology
6-Mar-2025
A new algorithm sheds light on ‘disordered’ #proteins
The #IntrinsicallyDisorderedProteins (IDPs) do not attain a stable secondary or tertiary structure and rapidly change their conformation, making #structurePrediction particularly challenging. These proteins although exhibit chaotic and ‘disordered’ structures, they still perform essential functions.
https://www.eurekalert.org/news-releases/1075837 #science
A new algorithm sheds light on ‘disordered’ proteins
Researchers from the Biomedical Sciences Research Center “Alexander Fleming” and the Centre for Misfolding Diseases at the University of Cambridge have found a way to predict the structure of a significant fraction of the proteome that was previously considered “dark” and difficult to be studied
Reliable analysis of intrinsically disordered proteins | Press and Public Relations
Reliable analysis of
#IntrinsicallyDisorderedProteins: Researchers of
#MainzUniversity and the European Molecular Biology Laboratory (EMBL) in Hamburg present a new approach to determine the form of disordered proteins by using two different methods simultaneously in a single sample 👉
https://press.uni-mainz.de/reliable-analysis-of-intrinsically-disordered-proteins/ #biophysics #MolecularBiology #SyntheticBiology #proteins #biocondensatesReliable analysis of intrinsically disordered proteins | Press and Public Relations
13-JUN-2024
Developing a system to study proteins without fixed structures
Researchers propose a novel pipeline that makes the analysis of intrinsically disordered proteins much faster and more convenient
I do love #IntrinsicallyDisorderedProteins but pinning them down to do a crystal structure is kind of against the spirit? NMR structures giving dynamic representations have served the field well
https://www.eurekalert.org/news-releases/1048185 #science #proteins #structuralBiology #IntrinsicDisorder

Developing a system to study proteins without fixed structures
Intrinsically disordered proteins (IDPs) can dynamically change their conformations depending on their external environment and can, therefore, bind to different compounds. However, they are difficult to analyze. Now, Tokyo Tech researchers have addressed this issue with a novel pipeline that enables a rapid crystal structure analysis of IDPs via a cell-free protein crystallization technique.

Scientists tame chaotic protein fueling 75% of cancers
<p>Meet MYC, the shapeless protein responsible for making the majority of human cancer cases worse. UC Riverside researchers have found a way to rein it in, offering hope for a new era of treatments. </p>