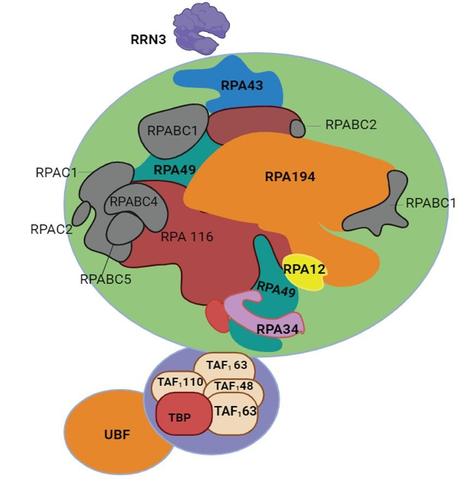
`Here, we show that the nascent polypeptide-associated complex controls the cotranslational modification of histones H2A and H4 by recruiting NatD and the upstream enzyme MetAP1 to ribosomes. MetAP1 and NatD cooperate on the ribosome to create a confined environment for the efficient sequential modification of the nascent histone chain.`
H2A.Z deposition by the SWR complex is stimulated by polyadenine DNA sequences in nucleosomes
The variant histone H2A.Z is deposited into nucleosomes immediately downstream of promoters, where it plays a critical role in transcription. The site-specific deposition of H2A.Z is catalyzed by the SWR complex, a conserved chromatin remodeler with affinity for promoter-proximal nucleosome-depleted regions (NDRs) and histone acetylation. By comparing the genomic distribution of H2A.Z in wild-type and SWR-deficient cells, we found that SWR is also responsible for depositing H2A.Z at thousands of non-canonical sites not directly linked to NDRs or histone acetylation. To understand the targeting mechanism of H2A.Z, we presented SWR to a library of canonical nucleosomes isolated from yeast and analyzed the preferred substrates. Our results revealed that SWR preferentially deposited H2A.Z into a subset of endogenous H2A.Z sites, which are overrepresented by polyadenine tracts on the top strands of the DNA duplex at the nucleosomal entry-exit sites. Insertion of polyadenine sequences into recombinant nucleosomes near the outgoing H2A-H2B dimer enhanced SWR’s affinity for the nucleosomal substrate and increased its H2A.Z insertion activity. These findings suggest that the genome encodes sequence-based information that facilitates remodeler-mediated targeting of H2A.Z.
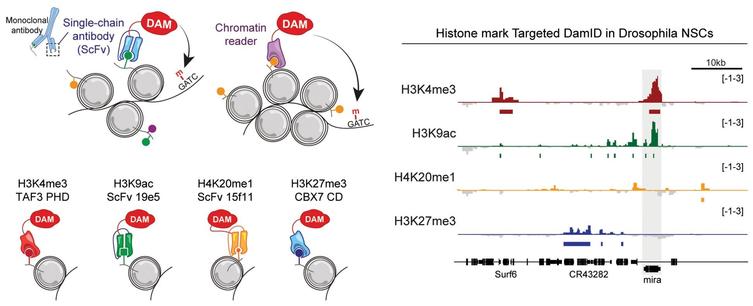
Targeted DamID detects cell-type-specific histone modifications in intact tissues or organisms
Current methods for profiling histone modifications require cell isolation and are not compatible with all tissue types. This study develops Targeted DamID, a method that can profile cell-type-specific histone marks throughout the genome, in vivo, in diverse model systems and without cell isolation.
Plant Histone Deacetylases: Their Classification and Inhibitor Search - Cytology and Genetics
Abstract Histone deacetylases is a family of enzymes pivotal in regulating numerous crucial cellular processes in both plant and animal cells. Plant histone deacetylases have been considerably less investigated in comparison to their human counterparts. This study aims to provide an in-depth characterization of histone deacetylases in two model plant proteins-species Arabidopsis thaliana and Oryza sativa. Phylogenetic analysis of their relationship to known human homologs has revealed their close relation to three classes of human histone deacetylases. Notably, the highest sequence homology among histone deacetylases from different evolutionary origins was observed between human HDAC6 and A. thaliana HDA5 (43.6% homology). Structural alignment results highlighted the conservation of catalytic domains and demonstrated a high activity of inhibitors against both histone deacetylases. Ligand-protein docking studies suggested a high potency of human histone deacetylase inhibitors against A. thaliana HDA5. These findings suggest the potential efficacy of human histone deacetylase inhibitors in modulating plant histone deacetylases, thereby enhancing growth regulation, development, and stress resistance in plants.
#Strawberry #ripening revealed: Key #histone modifications uncovered
#genomics #ChIP_seq #H3K9_K14ac #H3K27ac #H3K27me3
https://phys.org/news/2024-09-strawberry-ripening-revealed-key-histone.html
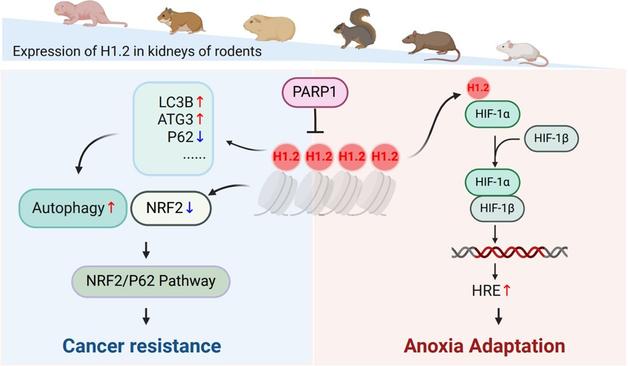
Comparative time-series multi-omics analyses suggest H1.2 involvement in anoxic adaptation and cancer resistance
Some long-lived animals, like the naked mole rat, are extraordinarily resistant to hypoxia and cancer. These authors show that embryonic fibroblasts from these animals overexpress histone 1.2, reducing tumor formation via the NRF2/P62 pathway and improving resistance to hypoxia.
dear #biology friends... I want to learn more about this:
"#Histone H4 is encoded in multiple genes at different loci including: HIST1H4A, HIST1H4B, HIST1H4C, HIST1H4D, HIST1H4E, HIST1H4F, HIST1H4G, HIST1H4H, HIST1H4I, HIST1H4J, HIST1H4K, HIST1H4L, HIST2H4A, HIST2H4B, HIST4H4."
Who can recommend a good read? I have questions like: are all genes expressed at the same time? Equally? How are all regulated? Do they all code the exact same 102 AAs? what about the biology of the up to 33 other AAs?
Organization Of DNA In Cell
#ArchaeaDNA, #DNA, #Histone, #Nucleosomes, #Supercoil #Genetics All living cells contain DNA in the form of double helix, its organization differs among cells in the three domains…. Medical Microbiology & Recombinant DNA Technology (RDT) Labs | Read More -
https://micrordt.wordpress.com/2024/04/27/organization-of-dna-in-cell/