Release Montseny Brook Newt · nf-core/proteinfold
[2.0.0] - 2026-03-27
Enhancements & fixes
[#177] - Fix typo in some instances of model preset alphafold2_ptm.
[PR #178] - Enable running multiple modes in parallel.
[#179] - Produce an interactive...
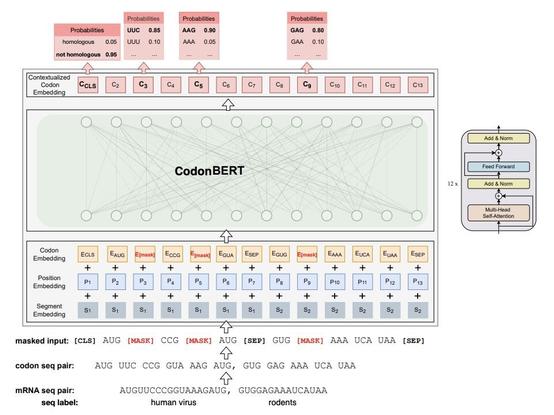
CodonBERT: Large Language Models for mRNA design and optimization
This is for those who think that #research is just writing paper in #academia and not in companies. Look how many are focused on AI software house partnership, and Sanofi even created #CondonBert, a language model that has been specifically trained to understand and generate #proteinsequences using the #Transformers architecture. It is an extension of #BERT https://www.biorxiv.org/content/10.1101/2023.09.09.556981v1
#LLM #machinelearning
Researchers Identify Blood Panel To Predict Placenta Accreta:
Of the nearly 4 million births each year in the United States, roughly 50,000 are marked by life-threatening complications, and up to 900 result in maternal death during delivery. One major, often life-threatening complication is placenta accreta spectrum (PAS), which poses a threat to both the mother and the baby.
https://www.nature.com/articles/s41598-022-24869-0
#ProteinSequences #proteins #molecularbiology #pregnancy #placenta #bloodwork #PAS

Circulating microparticle proteins predict pregnancies complicated by placenta accreta spectrum - Scientific Reports
Placenta accreta spectrum (PAS) is characterized by abnormal attachment of the placenta to the uterus, and attempts at placental delivery can lead to catastrophic maternal hemorrhage and death. Multidisciplinary delivery planning can significantly improve outcomes; however, current diagnostics are lacking as approximately half of pregnancies with PAS are undiagnosed prior to delivery. This is a nested case–control study of 35 cases and 70 controls with the primary objective of identifying circulating microparticle (CMP) protein panels that identify pregnancies complicated by PAS. Size exclusion chromatography and liquid chromatography with tandem mass spectrometry were used for CMP protein isolation and identification, respectively. A two-step iterative workflow was used to establish putative panels. Using plasma sampled at a median of 26 weeks’ gestation, five CMP proteins distinguished PAS from controls with a mean area under the curve (AUC) of 0.83. For a separate sample taken at a median of 35 weeks’ gestation, the mean AUC was 0.78. In the second trimester, canonical pathway analyses demonstrate over-representation of processes related to iron homeostasis and erythropoietin signaling. In the third trimester, these analyses revealed abnormal immune function. CMP proteins classify PAS well prior to delivery and have potential to significantly reduce maternal morbidity and mortality.
It's more or less common knowledge that most species that ever lived are now extinct. But it's less recognized that the same is true for
#ProteinSequences. For
#astrobiology, it's important to remember this when thinking about
#AncientLife 11/11