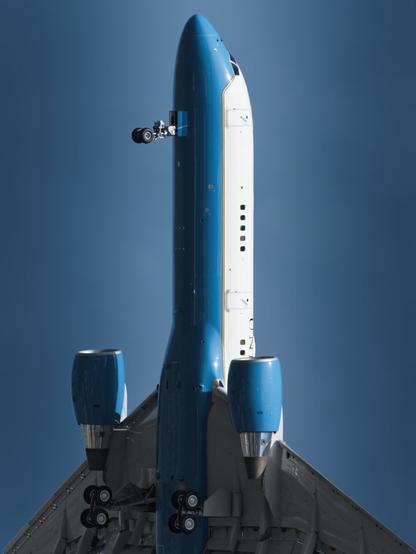
⬆️ Air Force 2
🛩️ Aircraft: 🇺🇸 Boeing C-32A military passenger transport aircraft
🌀 Powerplant: 2x 🇺🇸 Pratt & Whitney PW2000-40 high-bypass turbofan engines
🗂️ S/N: 99-0004
🧑✈️Piloted by: United States Air Force
📍Destination: Moffett Federal Airfield KNUQ, California, USA
🧭 Origin: Los Angeles International KLAX, California, USA
🖼️ Prints available 🔗
📷 Fujifilm X-H2
🔭 Fujinon XF500mmF5.6
.
.
.
.
#airforce
#usaf
#af2
#airforcetwo
#airforce2
#boeing
#c32
#knuq
#california
#usa
#planespotting
#aviation
#aviationphotography
#fujifilm
#fujixseries
🛩️ Aircraft: 🇺🇸 Boeing C-32A military passenger transport aircraft
🌀 Powerplant: 2x 🇺🇸 Pratt & Whitney PW2000-40 high-bypass turbofan engines
🗂️ S/N: 99-0004
🧑✈️Piloted by: United States Air Force
📍Destination: Moffett Federal Airfield KNUQ, California, USA
🧭 Origin: Los Angeles International KLAX, California, USA
🖼️ Prints available 🔗
📷 Fujifilm X-H2
🔭 Fujinon XF500mmF5.6
.
.
.
.
#airforce
#usaf
#af2
#airforcetwo
#airforce2
#boeing
#c32
#knuq
#california
#usa
#planespotting
#aviation
#aviationphotography
#fujifilm
#fujixseries