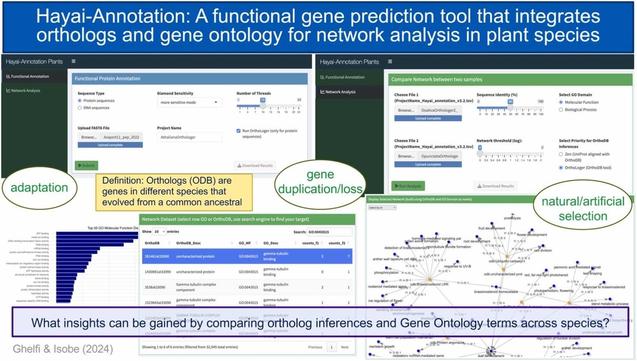
🐰🧬 The Easter bunny delivered early! The March 2026 Gene Ontology release is here with some impressive goodies:
📊 38,560 GO terms
🔬 8,883,119 annotations
🌍 5,557 species
💾 1,703,116 gene products
Your functional genomics just got a spring upgrade! Whether you're doing pathway analysis or comparative studies, this release has you covered.
🔗 current.geneontology.org
💬 https://community.alliancegenome.org/t/8648
#GeneOntology #Bioinformatics #OpenScience #FunctionalGenomics #Research #OpenData #AcademicMastodon

March 2026 GO Release now available
March 2026 GO Release (2026-03-25) Now Available We’re pleased to announce that the March 2026 Gene Ontology release is now live! Quick Access Links Current Release: https://current.geneontology.org Release Archive: https://release.geneontology.org DOI: Gene Ontology Data Archive Release Statistics Metric Count GO Terms 38,560 Annotations 8,883,119 Gene Products 1,703,116 Species 5,557 Detailed Release Information 📊 Complete statistics are available at the Gene Onto...