Noncanonical Structures in the Genome of Bovine Foamy Virus - Cytology and Genetics
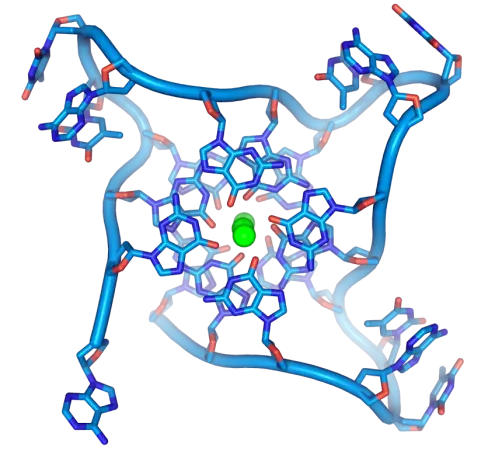
Abstract— Bioinformatic methods have been used to identify putative perfect G-quadruplexes (G4s) and three-way junctions (3WJs) in the bovine foamy virus (BFV) genome. Artificial intelligence (AI) AlphaFold 3 was used to confirm putative G4s and 3WJs by building 3D models of these noncanonical structures. G4s are secondary structures formed by G-rich sequences. Multihelical 3WJs formed by three duplexes connected at the binding point and G4s are considered as alternative structures in DNA and RNA that differ from the classical double-stranded B-DNA. In the present paper, the localization map of putative conservative intramolecular G4s formed by two G-tetrads on the BFV genome was created. Seven putative conservative G-quadruplexes in the sense strand of BFV proviral DNA and 22 G4s in the antisense strand formed by two G-tetrads with G-score from 32 to 36 were found by the multiple alignment of 37 BFV isolates with complete genome. The density of G4s was 0.6 G4/kb for the sense strand of the BFV proviral DNA, while it was 1.8 G4/kb for the antisense strand. One conservative 3WJ motif with a length of 73 bp with 100% homology localized in the 5'-untranslated region and partially on the 5'-end of the gag gene was found for a set of 37 BFV isolates. The 3WJ structure in BFV RNA is stabilized by 20 complementary bp with a free energy ΔG of 19.8 kcal/mol. The significance of this structure for BFV functioning has been proven. The use of AI AlphaFold 3 to build 3D models of theoretically determined perfect G4s and 3WJs in the BFV genome allowed for reliably determining putative alternative structures in nucleic acids.