#Bacterial species study challenges assumption that #structural similarity predicts #protein behavior
#Escherichia #Mycobacterium #cAMP #CRP #Allosteric #homology
https://phys.org/news/2025-02-bacterial-species-assumption-similarity-protein.html
Bacterial species study challenges assumption that structural similarity predicts protein behavior
A new study published in Biochemistry sheds light on how bacteria regulate their genes, challenging long-held assumptions about protein behavior. The research compares how two bacterial species—Escherichia coli and Mycobacterium tuberculosis—use a signaling molecule called cyclic AMP (cAMP) to control important cellular functions.
Explore unprecedented binding sites on protein targets by screening #covalent heterocyclic MiniFrags. See how they work identifying novel #allosteric sites on HDAC8 https://is.gd/CdPLih
Thanks @EnamineLtd
for making the covalent MiniFrags available https://is.gd/7LN1sb
"At the single-molecule level, we observe #stochastic sampling of eIF4F along the length of the messenger #RNA and identify #allosteric communication between the eIF4F subunits which ultimately drive cap-recognition and subsequent activation of the message. Our experiments uncover novel functions for each subunit of eIF4F and we conclude by presenting a model for messenger RNA activation which precisely defines the composition of the activated message."
"Here we use a range of reporter proteins to demonstrate that their #chimeras with duplicated #receptor domains produce YES gate protein switches with large (up to 9,000-fold) dynamic ranges and fast (minutes) response rates. In such switches, the #epistatic interactions between largely independent synthetic #allosteric sites result in an OFF state with minimal background noise."
Development of epistatic YES and AND protein logic gates and their assembly into signalling cascades - Nature Nanotechnology
Protein-based nanobiosensors with two epistatically interacting synthetic allosteric regulatory systems result in YES gate protein switches with large dynamic ranges and fast response times. These biosensors enable construction of rapid diagnostic tests compatible with clinical chemistry analysers.
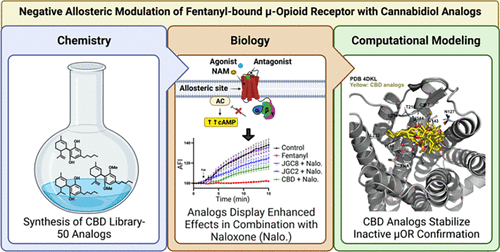
Structure–Activity Relationship #Study of #Cannabidiol-Based #Analogs as Negative #Allosteric #Modulators of the μ-#Opioid #Receptor
https://pubs.acs.org/doi/epdf/10.1021/acs.jmedchem.3c00061
“The US faces an unprecedented surge in fatal drug overdoses. Naloxone, the only antidote for opiate overdose, competes at the mu opioid receptor (μOR) orthosteric site. Naloxone struggles against fentanyl-class synthetic opioids that now cause ∼80% of deaths.”
Aravind Penmatsa and colleagues @iiscbangalore
https://www.embopress.org/doi/full/10.15252/embj.2023113418
Scientists develop strategy to engineer artificial #allosteric sites in #protein complexes.
https://phys.org/news/2023-07-scientists-strategy-artificial-allosteric-sites.html
Scientists develop strategy to engineer artificial allosteric sites in protein complexes
According to a recently published research paper, an approach has been developed to create artificial allosteric sites (where by binding an effector molecule, activity at the distal active site is regulated) in protein complexes. This breakthrough research holds significant promise for a wide range of applications in industrial, biological, medical, and agricultural fields. The work was published in Nature Chemistry.
Examining the role of 'blind dating' in #bacteria #evolution.
#OCP1 #FRP #protein #allosteric #regualtion #cyanobacterial #photoprotection
https://phys.org/news/2023-04-role-dating-bacteria-evolution.html
Examining the role of 'blind dating' in bacteria evolution
Proteins are the key players for virtually all molecular processes within the cell. To fulfill their diverse functions, they have to interact with other proteins. Such protein-protein interactions are mediated by highly complementary surfaces, which typically involve many amino acids that are positioned precisely to produce a tight, specific fit between two proteins. However, comparatively little is known about how such interactions are created during evolution.