Guide: Switching from Conda/Mamba to Pixi. Near instant environments, better HPC compatibility and reproducibility. But really, it's the speed!
Antonio Camargo
- 69 Followers
- 79 Following
- 131 Posts
Happy to announce a new preprint from the lab, together with fantastic @richardneher and Liam Shaw! 📝
https://www.biorxiv.org/content/10.1101/2024.07.08.602537v1
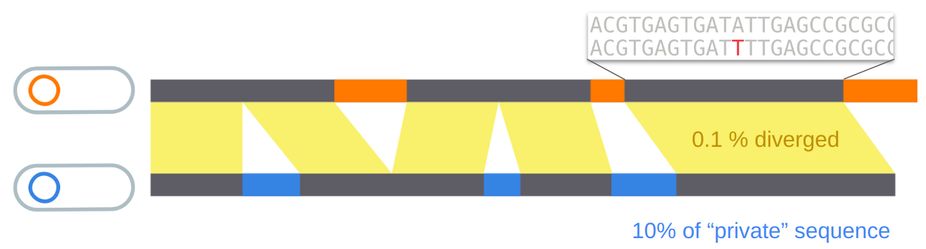
We were motivated by the following puzzling observation: microbial genomes can be extremely similar in the core genome, while still differing by large portions of accessory genome.
Therefore we ask the question:
1) how does this diversity accumulate?
2) at what rate do genomes undergo large structural changes?
A thread 🧵 [1/13]
GenomeSPOT: Genome-based Salinity, pH, Oxygen Tolerance, and Temperature for Bacteria and Archaea
GitHub: https://github.com/cultivarium/GenomeSPOT
Preprint: https://www.biorxiv.org/content/10.1101/2024.03.22.586313v1
Nice N&V Forum by @JakobWirbel, Ami Bhatt, Alexander Probst discussing how this study and the related Rodríguez del Río et al. provide insight into the vast structural and functional diversity of previously unknown microbial protein families
Preprint https://www.biorxiv.org/content/10.1101/2024.01.29.577392
Seen on the other place.
I have been mystified why K represented Lysine for many years. Undergrad #biochemistry lecturer said “I don't know why, but I assume it is because of the end of the side chain looks like a K”. Well, turns out the answer is more simple.
Source: M Eugenio Vazquez. Posted in honour of Margaret Dayhoff, inventor of the single-letter amino acid code