GitHub - dcjones/proseg: Probabilistic cell segmentation for in situ spatial transcriptomics
Probabilistic cell segmentation for in situ spatial transcriptomics - dcjones/proseg
🚨 Amazing work:
"Highly multiplexed spatial transcriptomics in bacteria" by Sarfatis et al. in Science
1000-fold volumetric expansion of E. coli cells paves the way to image-based single-cell transcriptomic 🦠 🔬
#MERFISH
https://www.science.org/doi/10.1126/science.adr0932
bioRxiv:
https://www.biorxiv.org/content/10.1101/2024.06.27.601034v1
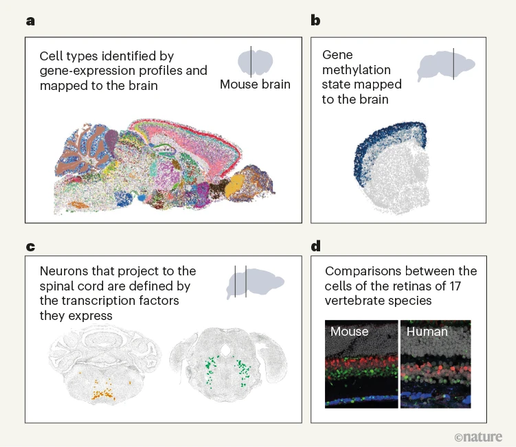
Maria Tosches & Heather Lee summarize the newly reported, and massive, atlases of mouse brain transcriptomics that include as well partial projectomes.
https://www.nature.com/articles/d41586-023-03781-1
A rather spectacular application of Xiaowei Zhuang's et al. 2015 #MERFISH spatial transcriptomics technology https://www.science.org/doi/10.1126/science.aaa6090
#neuroscience #mouse

Cellular atlases of the entire mouse brain
Transcriptomic and epigenomic data from millions of mouse brain cells.
COMMOT, COMMunication analysis by Optimal Transport, a versatile tool infer cell–cell communication in #SpatialTranscriptomics 🤠
Compatible with #MERFISH #Visium seqFISH+ #STARmap
vs #CellChat #Giotto #CellPhoneDBv3
Dr. Qing Nie lab Nature Methods 2023
https://www.nature.com/articles/s41592-022-01728-4

Screening cell–cell communication in spatial transcriptomics via collective optimal transport - Nature Methods
This work presents a computational framework, COMMOT, to spatially infer cell–cell communication from transcriptomics data based on a variant of optimal transport (OT).
If you work on the spatial transcriptome data. Our tool may help you
https://monkeybread.readthedocs.io/en/latest/ #merfish #spatialmonkeybread — monkeybread
Joint Sparse method for Imaging Transcriptomics (JSIT) for decoding imaging spatial transcriptomics data such as MERFISH. Better performance than the original pipeline (MERlin) according to the authors.
#SpatialTranscriptomics #matlab #merfish #bioinformatics #toolshttps://www.biorxiv.org/content/10.1101/2022.11.22.517523v1.fullJoint Sparse method for Imaging Transcriptomics (JSIT) for decoding imaging spatial transcriptomics data such as MERFISH. Better performance than the original pipeline (MERlin) according to the authors.
#SpatialTranscriptomics #matlab #merfish #bioinformatics #toolshttps://www.biorxiv.org/content/10.1101/2022.11.22.517523v1.full