Part 29 — Sometimes “smaller i...
Part 29 — Sometimes “smaller is elegant” in drug discovery. In early discovery, medicinal chemists often chase potency by making molecules larger and more lipophilic. Yet potency should always be… | Stan Van Boeckel
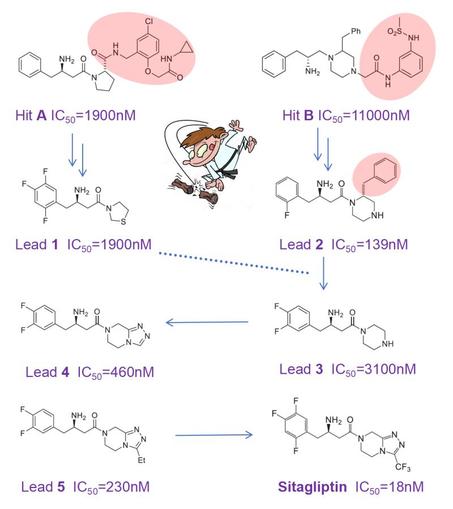
Part 29 — Sometimes “smaller is elegant” in drug discovery. In early discovery, medicinal chemists often chase potency by making molecules larger and more lipophilic. Yet potency should always be interpreted in the context of molecular weight and lipophilicity. Ligand‑efficiency metrics normalize potency to size and logP, helping chemists distinguish truly efficient hits and leads from bulky or greasy molecules that only appear potent but usually bring ADMET liabilities. The real challenge is to think broadly, so consider to make your hit or lead smaller during optimization, or 'just do it'. The discovery of sitagliptin illustrates this beautifully: a blockbuster drug that is smaller than the original screening hits. Sitagliptin (approved 2006) is the first DPP‑4 inhibitor that slows the rapid degradation of GLP‑1 and GIP, thereby lowering plasma glucose. In Merck’s HTS campaign, compounds A and B emerged as low‑potency hits. Here comes the twist: instead of growing these hits, the team chopped off large moieties and explored whether the resulting fragments could be optimized with small substituents such as fluorines. Surprisingly, this yielded compact leads 1 and 2 with improved potency, while selectivity across DPP family members and oral bioavailability in rat were continuously monitored. Lead 2 was then chopped again-taking inspiration from lead 1, which lacked its extra phenyl ring-to give weak lead 3. Accepting a 20‑fold drop in IC₅₀ seems counterintuitive, but it opened the door to expand the piperazine and create a new heterocycle. This delivered more potent lead 4, followed by lead 5. Further fluorination of 5 improved oral exposure and ultimately delivered the blockbuster sitagliptin. This small, well‑balanced molecule (MW 407; IC₅₀ 18 nM; oral bioavailability 87%; t½ 12 h; rapid absorption; logP 1.5; high selectivity; medium solubility; 5 rotatable bonds; HA/HD 4/1; non‑toxic) is a superb example of optimizing property space without molecular obesity. A remarkable achievement by Malcolm MacCoss, Ph.D., FRSC and his team at Merck.