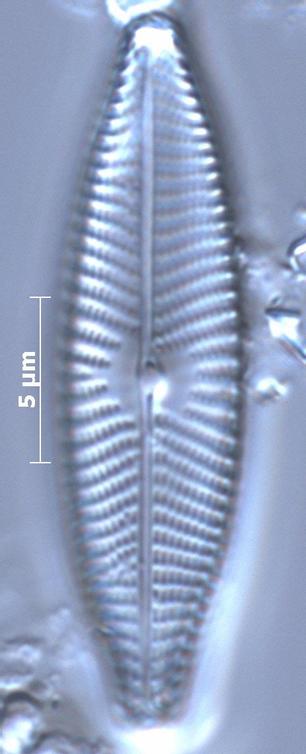
Gelis, Barry-Martinet, Briand, Chonova, Cochero, Gassiole, Kahlert, Karthick, Keck, Kelly, Kochoska, Mann, Nayak, Pfannkuchen, Servulo, Trobajo, Vasselon, Vidakovic, Viollaz, Wetzel, Zimmermann, Rimet, F. (2025). https://doi.org/10.1051/limn/2025009
https://carrtel-collection.hub.inrae.fr/barcoding-databases/phytool
#algae #bloom #nature #ecology #science #barcoding #metabarcoding #eDNA #omics
Gelis, Barry-Martinet, Briand, Chonova, Cochero, Gassiole, Kahlert, Karthick, Keck, Kelly, Kochoska, Mann, Nayak, Pfannkuchen, Servulo, Trobajo, Vasselon, Vidakovic, Viollaz, Wetzel, Zimmermann, Rimet, F. (2025). https://doi.org/10.1051/limn/2025009
Genetic Diversity of Ukrainian Populations of Invasive Species of the Genus Galinsoga Assessed by ISSR-Markers - Cytology and Genetics
Abstract Two species of the genus Galinsoga, G. parviflora Cav. and G. quadriradiata Ruiz and Pav., are among the most successful invasive plants causing significant damage to natural- and agroecosystems. Their natural distribution range extends from North to South America, and the adventitious part of the range includes all continents except Antarctica. Despite the practical importance of G. parviflora and G. quadriradiata, the genetic diversity of European populations of these species remains unexplored. In this study, ISSR markers were used to study Ukrainian populations of G. parviflora and G. quadriradiata and compared them with plants from Poland, Lithuania, and Portugal. The results obtained indicate the low genetic diversity (Shannon’s index I = 0.124) of G. quadriradiata populations, which is probably due to the small size of the original population introduced to the Old World from America. In contrast, the genetic diversity in G. parviflora populations is significantly higher (I = 0.254). Some genotypes of G. parviflora have a wide geographical distribution and, at the same time, different genotypes occur in the same area. The data obtained are in good agreement with the hypothesis that the main way of the invasion of Galinsoga species in the Old World was escape from botanical gardens. Among the samples examined, several forms of hybrid nature were identified, probably originating from hybrids between G. parviflora and G. quadriradiata, followed by subsequent backcrossing with one of the parent species.
Universal #RNA #barcoding system for tracking #gene_transfer in #bacteria created.
https://phys.org/news/2025-03-universal-rna-barcoding-tracking-gene.html
Universal RNA barcoding system for tracking gene transfer in bacteria created
In the microscopic world of bacteria, gene transfer is a powerful mechanism that can alter cellular function, drive antibiotic resistance and even shape entire ecosystems. Now an interdisciplinary group of researchers at Rice University has developed an innovative RNA "barcoding" method to track these genetic exchanges in microbial communities, providing new insights into how genes move across species. The findings were recently published in Nature Biotechnology.
Totally about 1K specimens in our collection have dna-derived data (attached sequences): https://www.gbif.org/dataset/d922b606-6c94-4d51-9277-36c9b03872a7
The Fungarium of Yugra State University
The Fungarium of Yugra State University is a systematic reference collection of fungi organized as a part of the Yugra State University Biological Collection (YSU BC). The main purpose of the Fungarium is to initiate and facilitate systematic studies of fungi in the region. It also serves for education and can be used by specialists in different applied disciplines.The taxonomical structure of the collections currently includes the total number of species represented – about 1.5K, genera represented – about 500, families represented – about 150. The majority of the specimens in the YSU Fungari…
20 years after Paul Hebert paper on DNA #barcoding who could expect this wordwide success?
#footprint #metabarcoding #algae #eDNA #barcoding #carbon
Excellent paper from F Keck.
Reference barcoding libraries problems:
(i) mislabelling, (ii) sequencing errors, (iii) sequence conflict, (iv) taxonomic conflict, (v) low taxonomic resolution, (vi) missing taxa and (vii) missing intraspecific variants. They give possible consequences on the taxonomic assignment process
Navigating the seven challenges of taxonomic reference databases in metabarcoding analyses
https://doi.org/10.1111/1755-0998.13746
#metabarcoding #DNA #bioinfo #barcoding