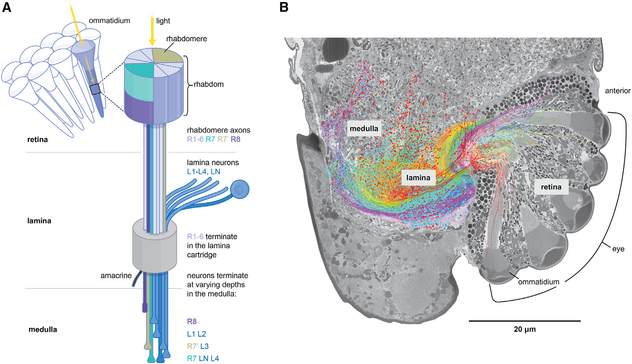
A connectome of the optic lobe of the extremely tiny fairy wasp, Megaphragma sp.
"A complete reconstruction of the early visual system of an adult insect", by Chua et al. 2023 (Chklovskii & Polilov) https://www.sciencedirect.com/science/article/pii/S096098222301237X
Don't miss the supplemental figures.
"Compared with the honeybee and the fruit fly, Megaphragma exhibits the following miniaturization-related adaptations: a significant reduction in the number of ommatidia, absence of several cell types, reduced size, and denucleation of neurons. Interestingly, the reduction in lens diameter is less than that expected from the optimization of the optical resolution of the eye. This suggests that light sensitivity is a more important
consideration when lens diameter approaches the wavelength of light. The absence of wide-field (or non-columnar) lamina neurons in Megaphragma could be a consequence of the smaller number of ommatidia, their larger acceptance angle, and the lower resolving power of the eye."
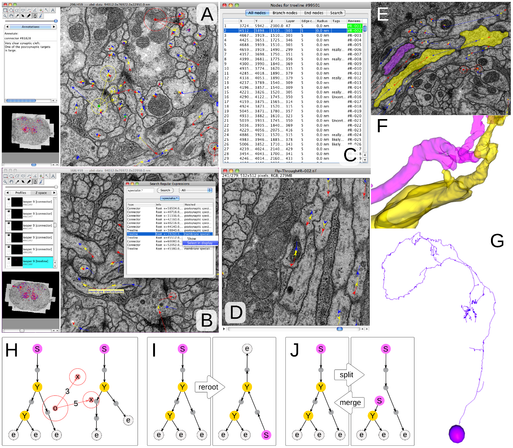
Volume assembled with #FijiSc and #TrakEM2, and its neurons and synapses mapped with #CATMAID. Woohoo!
#neuroscience #connectomics #VolumeEM #vEM #insects #miniaturization