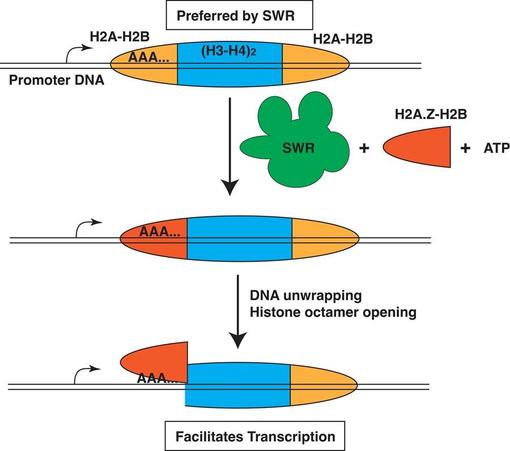
H2A.Z deposition by the SWR complex is stimulated by polyadenine DNA sequences in nucleosomes
The variant histone H2A.Z is deposited into nucleosomes immediately downstream of promoters, where it plays a critical role in transcription. The site-specific deposition of H2A.Z is catalyzed by the SWR complex, a conserved chromatin remodeler with affinity for promoter-proximal nucleosome-depleted regions (NDRs) and histone acetylation. By comparing the genomic distribution of H2A.Z in wild-type and SWR-deficient cells, we found that SWR is also responsible for depositing H2A.Z at thousands of non-canonical sites not directly linked to NDRs or histone acetylation. To understand the targeting mechanism of H2A.Z, we presented SWR to a library of canonical nucleosomes isolated from yeast and analyzed the preferred substrates. Our results revealed that SWR preferentially deposited H2A.Z into a subset of endogenous H2A.Z sites, which are overrepresented by polyadenine tracts on the top strands of the DNA duplex at the nucleosomal entry-exit sites. Insertion of polyadenine sequences into recombinant nucleosomes near the outgoing H2A-H2B dimer enhanced SWR’s affinity for the nucleosomal substrate and increased its H2A.Z insertion activity. These findings suggest that the genome encodes sequence-based information that facilitates remodeler-mediated targeting of H2A.Z.
#RNA #Transcription #RNApolymerase-II #Histones
https://www.science.org/doi/10.1126/sciadv.adt3037
Researchers succeed at generating 3D #visualizations of #chloroplasts' copying machines.
https://phys.org/news/2024-03-succeed-generating-3d-visualizations-chloroplasts.html
Researchers succeed at generating 3D visualizations of chloroplasts' copying machines
For life on Earth, it is essential that plants carry out photosynthesis and ultimately produce oxygen and chemical energy with the help of sunlight. Researchers from Göttingen and Hannover have now succeeded for the first time in visualizing the copying machine of chloroplasts, the RNA polymerase PEP, in high-resolution 3D.
#cellbiology #transcription #RNAPolymerase #geneexpression
The RNA export and RNA decay complexes THO and TRAMP prevent transcription-replication conflicts, DNA breaks, and CAG repeat contractions
Expansion of structure-forming CAG/CTG repetitive sequences is the cause of several neurodegenerative disorders and deletion of repeats is a potential therapeutic strategy. Transcription-associated mechanisms are known to cause CAG repeat instability. In this study, we discovered that Thp2, an RNA export factor and member of the THO (suppressors of transcriptional defects of hpr1Δ by overexpression) complex, and Trf4, a key component of the TRAMP (Trf4/5-Air1/2-Mtr4 polyadenylation) complex involved in nuclear RNA polyadenylation and degradation, are necessary to prevent CAG fragility and repeat contractions in a Saccharomyces cerevisiae model system. Depletion of both Thp2 and Trf4 proteins causes a highly synergistic increase in CAG repeat fragility, indicating a complementary role of the THO and TRAMP complexes in preventing genome instability. Loss of either Thp2 or Trf4 causes an increase in RNA polymerase stalling at the CAG repeats and other genomic loci, as well as genome-wide transcription-replication conflicts (TRCs), implicating TRCs as a cause of CAG fragility and instability in their absence. Analysis of the effect of RNase H1 overexpression on CAG fragility, RNAPII stalling, and TRCs suggests that RNAPII stalling with associated R-loops are the main cause of CAG fragility in the thp2Δ mutants. In contrast, CAG fragility and TRCs in the trf4Δ mutant can be compensated for by RPA overexpression, suggesting that excess unprocessed RNA in TRAMP4 mutants leads to reduced RPA availability and high levels of TRCs. Our results show the importance of RNA surveillance pathways in preventing RNAPII stalling, TRCs, and DNA breaks, and show that RNA export and RNA decay factors work collaboratively to maintain genome stability.
The RNA export and RNA decay complexes THO and TRAMP prevent transcription-replication conflicts, DNA breaks, and CAG repeat contractions
Expansion of structure-forming CAG/CTG repetitive sequences is the cause of several neurodegenerative disorders and deletion of repeats is a potential therapeutic strategy. Transcription-associated mechanisms are known to cause CAG repeat instability. In this study, we discovered that Thp2, an RNA export factor and member of the THO (suppressors of transcriptional defects of hpr1Δ by overexpression) complex, and Trf4, a key component of the TRAMP (Trf4/5-Air1/2-Mtr4 polyadenylation) complex involved in nuclear RNA polyadenylation and degradation, are necessary to prevent CAG fragility and repeat contractions in a Saccharomyces cerevisiae model system. Depletion of both Thp2 and Trf4 proteins causes a highly synergistic increase in CAG repeat fragility, indicating a complementary role of the THO and TRAMP complexes in preventing genome instability. Loss of either Thp2 or Trf4 causes an increase in RNA polymerase stalling at the CAG repeats and other genomic loci, as well as genome-wide transcription-replication conflicts (TRCs), implicating TRCs as a cause of CAG fragility and instability in their absence. Analysis of the effect of RNase H1 overexpression on CAG fragility, RNAPII stalling, and TRCs suggests that RNAPII stalling with associated R-loops are the main cause of CAG fragility in the thp2Δ mutants. In contrast, CAG fragility and TRCs in the trf4Δ mutant can be compensated for by RPA overexpression, suggesting that excess unprocessed RNA in TRAMP4 mutants leads to reduced RPA availability and high levels of TRCs. Our results show the importance of RNA surveillance pathways in preventing RNAPII stalling, TRCs, and DNA breaks, and show that RNA export and RNA decay factors work collaboratively to maintain genome stability.
The RNA export and RNA decay complexes THO and TRAMP prevent transcription-replication conflicts, DNA breaks, and CAG repeat contractions
Expansion of structure-forming CAG/CTG repetitive sequences is the cause of several neurodegenerative disorders and deletion of repeats is a potential therapeutic strategy. Transcription-associated mechanisms are known to cause CAG repeat instability. In this study, we discovered that Thp2, an RNA export factor and member of the THO (suppressors of transcriptional defects of hpr1Δ by overexpression) complex, and Trf4, a key component of the TRAMP (Trf4/5-Air1/2-Mtr4 polyadenylation) complex involved in nuclear RNA polyadenylation and degradation, are necessary to prevent CAG fragility and repeat contractions in a Saccharomyces cerevisiae model system. Depletion of both Thp2 and Trf4 proteins causes a highly synergistic increase in CAG repeat fragility, indicating a complementary role of the THO and TRAMP complexes in preventing genome instability. Loss of either Thp2 or Trf4 causes an increase in RNA polymerase stalling at the CAG repeats and other genomic loci, as well as genome-wide transcription-replication conflicts (TRCs), implicating TRCs as a cause of CAG fragility and instability in their absence. Analysis of the effect of RNase H1 overexpression on CAG fragility, RNAPII stalling, and TRCs suggests that RNAPII stalling with associated R-loops are the main cause of CAG fragility in the thp2Δ mutants. In contrast, CAG fragility and TRCs in the trf4Δ mutant can be compensated for by RPA overexpression, suggesting that excess unprocessed RNA in TRAMP4 mutants leads to reduced RPA availability and high levels of TRCs. Our results show the importance of RNA surveillance pathways in preventing RNAPII stalling, TRCs, and DNA breaks, and show that RNA export and RNA decay factors work collaboratively to maintain genome stability.