📰 "Mathematical Modeling of Early Embryonic Cell Cycles of Drosophila melanogaster"
https://arxiv.org/abs/2605.06598 #DrosophilaMelanogaster #Drosophila #Behaviour #Q-Bio.Cb
#Q-Bio.Qm
#Embryo
Mathematical Modeling of Early Embryonic Cell Cycles of Drosophila melanogaster
In the early stages of development, Drosophila melanogaster embryos possess very fast and well-coordinated cell cycles. In the cell cycle, CDK activity is essentially regulated by binding CDK and CycB to form an active complex and by phosphorylating CDK via CDC25 and dephosphorylating it via Wee1. We develop a mathematical model for the embryonic cell cycle which is biochemically sound and which can be rigorously analysed after a model reduction. We show that there exists a region in the parameter space where the model describes oscillations. We then focus on the role of two parameters: the CycB synthesis and the activation coefficient of APC. Our main biological hypothesis is that the first one is responsible for the period lengthening over the first 14 cycles which can be experimentally observed and this hypothesis is supported by numerical simulations of our model: if the CycB synthesis is made time-dependent with a prescribed dynamics, then our simulations show qualitatively a very similar behavior to experimental data reported in the literature.

NASP functions in the cytoplasm to prevent histone H3 aggregation during early embryogenesis
Das and Chavez et al. demonstrate that in D. melanogaster, the histone chaperone NASP indirectly affects the nuclear import and chromatin deposition of H3.
📰 "Bitesize bundles F-actin and influences actin remodeling in syncytial Drosophila embryo development"
https://doi.org/doi:10.1083/jcb.202306071https://pubmed.ncbi.nlm.nih.gov/42089875/ #Drosophila #Embryo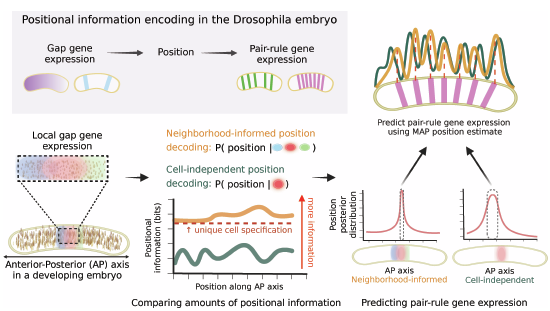
Neighborhood-informed positional information for precise cell identity specification - Molecular Systems Biology
During development, cells reliably establish their identities, a process that is enabled in part by positional information encoded in gene expression patterns. Previous works showed that cells in Drosophila embryos can utilize this information to decode their position along the anterior-posterior axis with a 1% embryo-length positional precision. However, this precision is insufficient to uniquely determine position, leading to a positional information gap. Here, we propose a neighborhood-informed information-theoretic framework which allows to quantitatively estimate the amount of information regarding position which exists in the microenvironment of each cell. We formulate how much additional information exists in neighboring cells as a function of spatial variation in gene expression. We show that the additional information encoded by local neighborhoods is sufficient to uniquely specify cell identities, closing the information gap on average across major patterning axes in Drosophila embryos, gastruloids, and the developing neural tube. Furthermore, neighborhood-informed decoders predict cell positions and downstream gene expression patterns more accurately than cell-independent decoders, resulting in lower decoding variability, which is maintained in mutant Drosophila embryos. Our results provide a basis for the analysis of cellular decision-making in the context of their microenvironments.

LGL-1 and the RhoGAP protein PAC-1 redundantly polarize the Caenorhabditis elegans embryonic epidermis
Apical–basal polarity is regulated by conserved cortical proteins, but their requirements vary between organisms. Here, we show that LGL-1 and the RhoGAP P
📰 "Synergy of RNA Concentration, RNA Binding Proteins, and RNA Palindrome Drives clustering of oskar mRNA in vivo"
https://www.biorxiv.org/content/10.64898/2026.04.30.721893v1?rss=1 #Drosophila #Embryo📰 "H2A.Z levels control the timing of major events at the maternal-zygotic transition"
https://www.biorxiv.org/content/10.64898/2026.04.27.721027v1?rss=1 #Drosophila #Embryo


