RE: https://social.edu.nl/@intconfchemstr/116416143010752584
Great to see this paper finally come out. We developed a generative model, ANNalog, that takes a SMILES string and generates medchem analogs. This was trained using pairs of molecules from #chembl. More on my blog: https://baoilleach.blogspot.com/2026/04/annalog-generative-model-for-medchem.html
New Blog Post: Prioritizing Drug-Like 💊 ChEMBL Compounds Within Target 🎯 Profiles
In this post, I go through how to use the #Python #ChEMBL #API and #SQLite to:
• Retrieve compound and target activity data programmatically
• Build a local database of molecules and their associated targets
• Rank compounds based on Lipinski Rule of Five violations
Read it at https://bertiewooster.github.io/2026/01/05/ChEBML-database.html. Marimo and Jupyter notebooks too!
#cheminformatics #drugDiscovery #chemistry #medChem #medicinalChemistry
Prioritizing Drug-Like ChEMBL Compounds Within Target Profiles
When reviewing data to find pharma compounds for virtual screening, we might want to check what their target profiles and rank candidates by how many Lipinski’s rule of five violations they have–the fewer the better. Here, a target profile refers to the set of targets a compound is known to be active against. This post uses the ChEMBL API and a SQLite database to do that.
ChEMBL 36 is out! if you're using chembl-downloader for all of your ChEMBL needs, then you just have to re-run your reproducible workflows and get arbitrarily new and better results, you beautiful nerd
https://chembl.blogspot.com/2025/09/chembl-36-is-out.html
#chembl #cheminformatics #reproducibility

ChEMBL 36 is out!
📊 Database Scale 🆕 New Data Sources Src_ID 72 – Chemical Probe data from Scientific Literature NLP and manual extraction from probe-r...
I used chembl-downloader to create some nice charts on how the number of compounds, assays, activities, and other entities in ChEMBL have grown over time
📖 https://cthoyt.com/2025/08/26/chembl-history.html
#chembl #chemistry #chemometrics #chemoinformatics #cheminformatics #rdkit #cdk #proteochemometrics
A historical analysis of ChEMBL
I’ve recently submitted an article to the Journal of Open Source Software (JOSS) describing chembl-downloader, a Python package for automating downloading and using ChEMBL data in a reproducible way. In this post, I use chembl-downloader to show how the number of compounds, assays, activities, and other entities in ChEMBL have changed over time.
new blog: "One Million IUPAC names #4: a lot is happening" https://chem-bla-ics.linkedchemistry.info/2025/08/09/one-million-iupac-names-4.html
"A lot is happening. If you have been following this project more closesly, you may have already seen some interesting updates, but I will post it here too."
replies to this post become blog comments.
#iupac #chemistry #openscience #chembl #beilstein
One Million IUPAC names #4: a lot is happening
A lot is happening. If you have been following this project more closesly, you may have already seen some interesting updates, but I will post it here too. First, a quick recap. In March I started a new Blue Obelisk project to collect CCZero IUPAC names from primary literature (paper still pending). It turned out we can automate that, while legally not violating any laws or licenses. In April I reported on some tweaks boosting the efficiency of the use of the API. I also reported on some possible further steps, including how to use the extracted names to create a larger set. Indeed, in June I could report to have passed the 200k IUPAC names, which with the idea from April gave us more than 1M IUPAC names.
Most cheminformatics code that queries ChEMBL struggles with reproducibility.
chembl-downloader can help:
>>> import chembl_downloader as cd
>>> df = cd.query("""
SELECT chembl_id, pref_name
FROM molecule_dictionary
WHERE pref_name IS NOT NULL
""")
It's even sneaking its way into @wpwalters and @dr_greg_landrum blogs :)
Code/Docs: https://github.com/cthoyt/chembl-downloader
Preprint: https://arxiv.org/pdf/2507.17783
#cheminformatics #chemoinformatics #chembl #reproducibility #chemistry #openscience

GitHub - cthoyt/chembl-downloader: Write reproducible code for getting and processing ChEMBL
Write reproducible code for getting and processing ChEMBL - cthoyt/chembl-downloader
Home - Building a simple deep learning model about adverse drug reactions
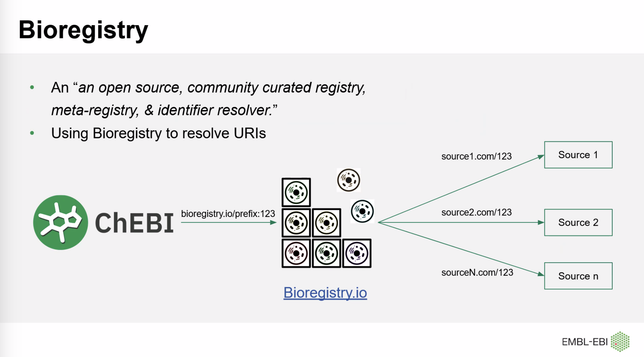
At the ChEBI 2.0 workshop, Muhammad Arsalan is presenting how ChEBI is using the Bioregistry to standardize its cross-references, generate URLs on their front-end, and more
#chembl #ebi #chebi #sssom #cheminformatics
An update on an older post looking at saving a relatively large csv file (although may not be considered large by some) as a Parquet file first (to be followed by 3 other smaller posts later detailing the use of Polars with scikit-learn without using Pandas at all)
https://jhylin.github.io/Data_in_life_blog/posts/21_ML1-1-1_Small_mols_in_chembl_update/ML1-1-1_chembl_cpds_parquet_new.html
#Scikit_Learn #Polars #parquet #Python #ChEMBL #Cheminformatics
Home - Small molecules in ChEMBL database
Here are some snapshots from the #ChEMBL symposium! Dr. Samantha attended & delivered a wonderful talk about the #SemanticWeb! You can find the slides right here ➡️ https://zenodo.org/records/13882075
#PSDI #event #semantics #semanticweb
The Semantic Web is Dead – Long Live the Semantic Web The Future of Semantics in the Physical Sciences (and beyond!)
Presentation by Dr Samantha Pearman-Kanza on The Semantic Web is Dead – Long Live the Semantic Web The Future of Semantics in the Physical Sciences (and beyond!) for the ChEMBL 15 Year Symposium. This presentation gave a hollistic overview of the Semantic Web, discussing common misconceptions, barriers and challenges, mitigations and suggestions for best practice, and a commentary on emerging use cases for semantics for the physical sciences.