If you've ever done a bit of #ProteinEngineering or #DirectedEvolution you know that funny things can happen 🤯 – and sometimes it's worth following up!
I'm excited to share that our paper is finally out in Nature Chemistry! 🎉
Read it here:
https://www.nature.com/articles/s41557-024-01490-4
Selection of a promiscuous minimalist cAMP phosphodiesterase from a library of de novo designed proteins - Nature Chemistry
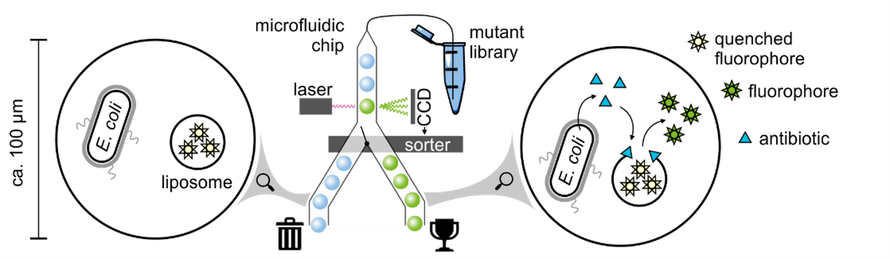
Evolution separates complex modern enzymes from their hypothetical simpler early ancestors, which raises the question of how unevolved sequences can develop new functions. Here a library of non-natural protein sequences was subjected to ultrahigh-throughput screens in microfluidic droplets, leading to the isolation of a phosphodiesterase enzyme capable of hydrolysing the biological second messenger, cyclic AMP.