#MetabolicPathway
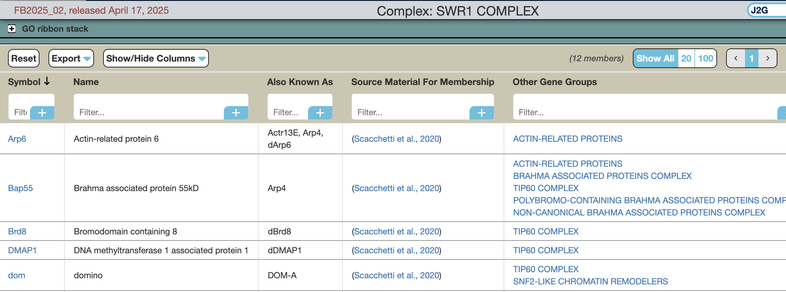
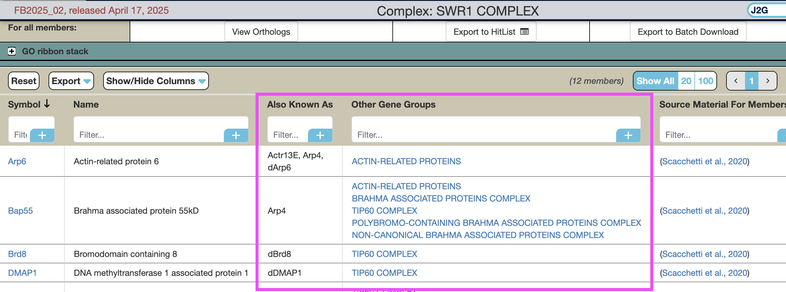
As with other interactive tables, users can customize the 'Members' table by reordering or filtering columns, or filtering which columns to display.
#FlyBaseTootorial
Referenced link: https://phys.org/news/2023-06-scientists-quantify-factors-contributing-flux.html
Discuss on https://discu.eu/q/https://phys.org/news/2023-06-scientists-quantify-factors-contributing-flux.html
Originally posted by Phys.org / @physorg_com: http://nitter.platypush.tech/physorg_com/status/1671147192562614273#m
Scientists quantify regulation factors contributing to #flux changes in the central #metabolicpathway in yeast @PNASNews https://pnas.org/doi/10.1073/pnas.2302779120 https://phys.org/news/2023-06-scientists-quantify-factors-contributing-flux.html
Scientists quantify regulation factors contributing to flux changes in the central metabolic pathway in yeast
Metabolic reaction flux change is the final result of interacting regulations by intracellular gene expression, transcriptional regulation, protein level, translation modification, and allosteric effect. However, the regulatory mechanism of metabolic flux changes is still not very clear.