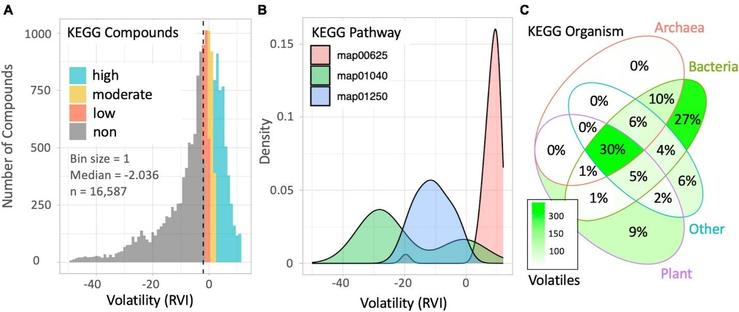
Calculated volatilities for all metabolites and their related genomes in the KEGG database
Volatility describes the tendency for a compound to partition into the gas phase and volatile metabolites facilitate unique biological interactions which have an influence on Earth's atmospheric physics and chemistry. Estimating which metabolites may be volatile is difficult, especially for those which do not have measured vapor pressures. Volcalc is a newly developed vapor pressure estimation tool which utilizes the SIMPOL.1 method, allowing users to rapidly identify plausible volatile metabolites within the Kyoto Encyclopedia for Genes and Genomes (KEGG) database. Here, we estimate the volatiles of all KEGG metabolites and associate them with KEGG reactions, enzymes, orthologs (KOs) and whole genomes within the KEGG database. This information may be used to identify which genes or species may be linked to particular forms of volatile metabolism, for the purpose hypothesis generation and integration into additional bioinformatics pipelines. This data is listed as a compliment to the publication "Automating methods for estimating metabolite volatility". The column "Paper" indicates whether the listed species is one from the subset analyzed within the data for Figure 3.For inquiries regarding the contents of this dataset, please contact the Corresponding Author listed in the README.txt file. Administrative inquiries (e.g., removal requests, trouble downloading, etc.) can be directed to [email protected]
Scientists uncover a multibillion-year epic written into the #chemistry_of_life.
#purine #biosynthesis #KEGG #evolution #ATP
https://phys.org/news/2024-05-scientists-uncover-multibillion-year-epic.html
Scientists uncover a multibillion-year epic written into the chemistry of life
The origin of life on Earth has long been a mystery that has eluded scientists. A key question is how much of the history of life on Earth is lost to time. It is quite common for a single species to "phase out" using a biochemical reaction, and if this happens across enough species, such reactions could effectively be "forgotten" by life on Earth.
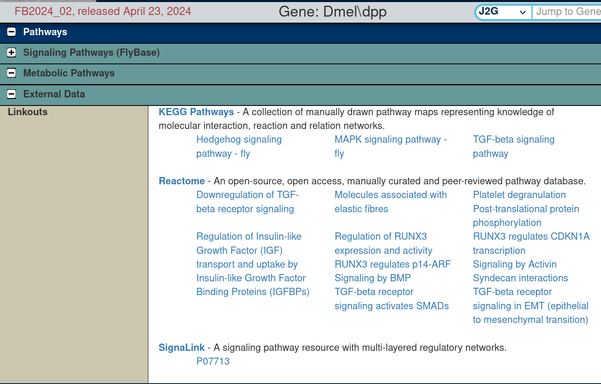
We've refreshed the links between FlyBase genes and #KEGG https://www.kegg.jp/kegg/pathway.html and #Reactome https://reactome.org/pathways. These links are displayed in the Pathways section (External Data subsection) of FlyBase gene reports and display the name of the KEGG/Reactome pathway(s) associated with the given gene at those databases.
#ggkegg: An #R package to analyze & visualize #KEGG information with #ggplot2:
Article: https://doi.org/10.1093/bioinformatics/btad622
Docu: https://noriakis.github.io/software/ggkegg/
GitHub: https://github.com/noriakis/ggkegg
#DIYbio #bioinformatics #CompChem #CitizenPharma #OpenAccess #OpenSource
Jouer avec l'API de KEGG | bioinfo-fr.net
Il n'est pas rare que nous ayons un jour besoin de récupérer des informations de la base de données KEGG (Kyoto Encyclopedia of Genes and Genomes). Cette base de données fournit un nombre conséquent d'informations sur les génomes et les réseaux de gènes mais également sur les voies métaboliques ou les maladies. Dans ces cas là, bien souvent, nous passons directement par le site internet à l'adresse http://www... Lire la suite...