HOST DEFENSE PEPTIDES https://flybase.org/reports/FBgg0002156, NUCLEAR EXOSOME TARGETING COMPLEX https://flybase.org/reports/FBgg0002165 and MITOCHONDRIAL INTERMEMBRANE SPACE TIM CHAPERONE COMPLEXES https://flybase.org/reports/FBgg0002014.
HOST DEFENSE PEPTIDES https://flybase.org/reports/FBgg0002156, NUCLEAR EXOSOME TARGETING COMPLEX https://flybase.org/reports/FBgg0002165 and MITOCHONDRIAL INTERMEMBRANE SPACE TIM CHAPERONE COMPLEXES https://flybase.org/reports/FBgg0002014.
And some new family gene groups: SPECTRINS https://flybase.org/reports/FBgg0002154 and SINGLE VON WILLEBRAND FACTOR TYPE C DOMAIN PROTEINS https://flybase.org/reports/FBgg0002153
#GeneGroups
and a GO-Causal Activity Model (GO-CAM https://geneontology.org/docs/gocam-overview/) that is based directly on GO annotation data.
Useful links to external resources (FlyCyc, Reactome, KEGG) and publications are provided, as well as options to export the gene list to analysis, download and orthology tools.
For more information, see this Commentary: http://flybase.org/commentaries/2024_12/MetabolicPathways.html
New #GeneGroups subset: Metabolic Pathways!
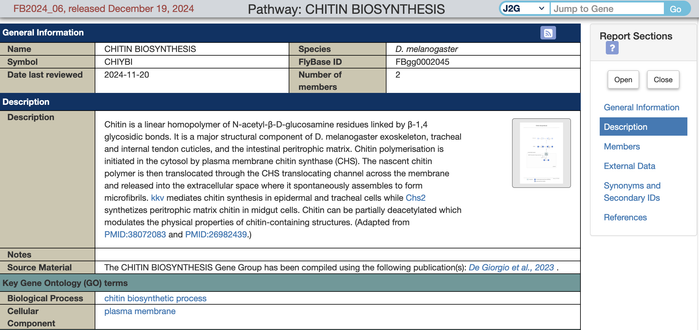
These manually curated reports aim to facilitate metabolic research in Drosophila by providing high-quality gene lists and visual representations of fly metabolic pathways, integrated with the other rich data in FlyBase. For example, see CHITIN BIOSYNTHESIS http://flybase.org/reports/FBgg0002045.htm
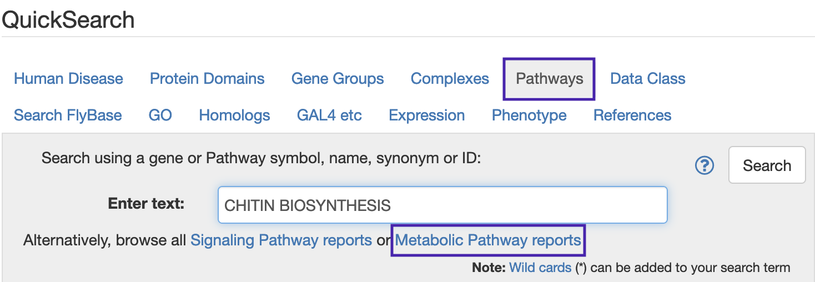
You can access Metabolic Pathways by searching via the Pathways QuickSearch tab, or by browsing the Metabolic Pathways list http://flybase.org/lists/FBgg/metabolicpathways
We've added >30 new reports to the Gene Group collection, many of which are enzyme complexes and functional groups related to metabolic processes and mitochondrial function. These include the GLUTAMYL-TRNA(GLN) AMIDOTRANSFERASE COMPLEX https://flybase.org/reports/FBgg0002000.htm, SERINE C-PALMITOYLTRANSFERASE COMPLEX https://flybase.org/reports/FBgg0002007.htm, UDP-N-ACETYLGLUCOSAMINE TRANSFERASE COMPLEX https://flybase.org/reports/FBgg0002009.htm, GAMMA-BUTYROBETAINE HYDROXYLASES https://flybase.org/reports/FBgg0002012.htm,
#FlyBaseTootorial #MacromolecularComplexes #GeneGroups
New in #GeneGroups! Have you wanted a one-stop resource to find Drosophila genes involved in transmembrane transport? We've compiled the Transmembrane Transporter Collection.
TRANSMEMBRANE TRANSPORTERS http://flybase.org/reports/FBgg0001934 was assembled from multiple sources, including data from Gene Ontology (GO) annotation, the Transporter Classification Database https://tcdb.org/ and publications. (we've reviewed and updated GO annotation to be more accurate and to provide better coverage). @go