Global Nature Positive Summit - October 2024 🌱
https://www.youtube.com/watch?v=Wl2b649IrDU
Her Uncle #DonCheetoMussolini 's #Tariffs Destroy Own Party-#GNPs BEG for Mercy
KARMA: Trump’s Tariffs DESTROY Own Party—Republicans BEG for Mercy!
More photo's from the Global Nature Positive Summit 🌿
#GNPS #kessmedia #sydney #animals #nature #commucation #iccsydney
Daniel Mookhey, Treasurer of New South Wales speaking at the Global Nature Positive Summit 🌿
#kessmedia #danielmookhey #GNPS #speaker #nsw #podium #sydneyaustralia
Photo of Tanya Plibersek speaking at the recent Global Nature Summit. She is the Minister for the Environment and Water of Australia 🌱 💧
#tanyaplibersek #GNPS #kessmedia #speaker #minister #podium #australia #environment
The Global Nature Positive Summit was held on October 8-10, 2024, on Gadigal Country, Sydney, Australia.
The Summit was a significant breakthrough in the global movement toward establishing nature positive economies and accelerating collective action to drive investment in nature. 🌿 🐳 🌱
#GNPS #kessmedia #naturepositive #sydney #global #australia #gadigalcountry #globalmovement #pannel #collabration
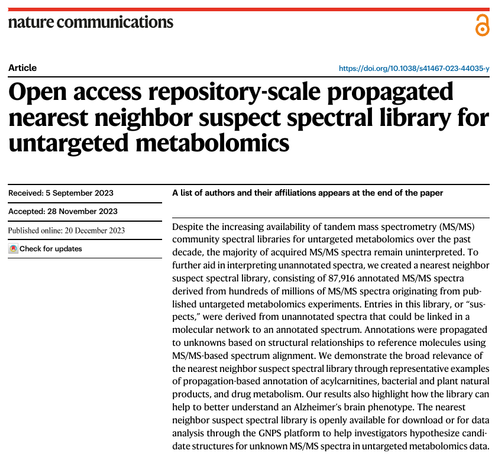
⚡ I'm super excited that our work on creating the #GNPS nearest neighbor suspect spectral library for #untargeted #metabolomics has now officially been published in @naturecomms.
This was a massive effort, as evidenced by the many awesome co-authors I was able to collaborate with. 🙌 Many thanks to everyone, but especially to @pdorrestein1, as this paper is one of the main outcomes from my postdoc at @ucsandiego.
Manuscript: https://doi.org/10.1038/s41467-023-44035-y
ENPKG integrates or is built on many computational metabolomics tools, such as #LOTUS, #SIRIUS, #GNPS, #matchms, #spec2vec, #GNPSDashboard, or #MassQL! A big thank you to the people behind them 🙏
➡ More info in the preprint: https://doi.org/10.26434/chemrxiv-2023-sljbt
A Sample-Centric and Knowledge-Driven Computational Framework for Natural Products Drug Discovery
Modern natural products (NPs) research relies on untargeted liquid chromatography coupled with mass spectrometry metabolomics. Together with cutting-edge processing and computational annotation strategies, such approaches can yield extensive spectral and structural information. However, current processing workflows require feature-alignment steps based on retention time which hinders the comparison of samples originating from different batches or analyzed using different instrumental setups. In addition, there is currently no analytical framework available to efficiently match processed metabolomics data and associated metadata with external resources. To address these limitations, we present a new sample-centric and knowledge-driven framework allowing multi-modal data alignment - e.g. through chemical structures, biological activities, or spectral features - and demonstrate its value in exploring large and chemodiverse natural extract datasets. Here, the experimental data is processed at the sample level, matched with external identifiers where possible, semantically enriched, and integrated into a unified knowledge graph. The use of semantic web technology enables comparison of processed and standardized data, information, and knowledge at the repository scale. We demonstrate the utility of the developed framework, the Experimental Natural Products Knowledge Graph (ENPKG), to leverage the results obtained from screening 1,600 plant extracts against trypanosomatids and streamline the identification of new antiparasitic compounds. Thanks to its versatility, the proposed approach allows for a radically novel exploitation of metabolomics data. Semantic web technologies are a fundamental asset and we anticipate that their adoption will complement the current computational metabolomics pipelines and enable the community to advance in the description of global chemodiversity and drug discovery projects.
MS/MS-Based Molecular Networking: An Efficient Approach for Natural Products Dereplication https://www.mdpi.com/1420-3049/28/1/157#
#metabolomics #GNPS #molecularnetworking #MSMS #spectra #LCMS
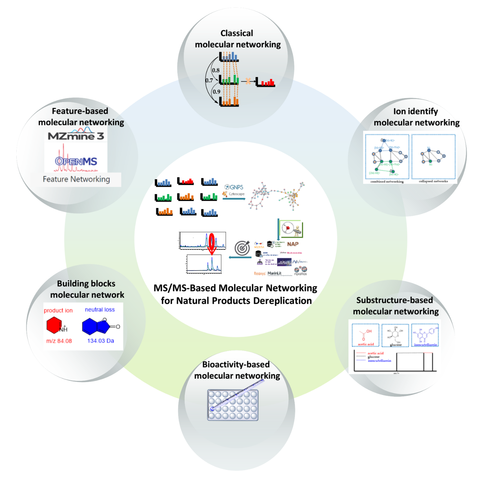
MS/MS-Based Molecular Networking: An Efficient Approach for Natural Products Dereplication
Natural products (NPs) have historically played a primary role in the discovery of small-molecule drugs. However, due to the advent of other methodologies and the drawbacks of NPs, the pharmaceutical industry has largely declined in interest regarding the screening of new drugs from NPs since 2000. There are many technical bottlenecks to quickly obtaining new bioactive NPs on a large scale, which has made NP-based drug discovery very time-consuming, and the first thorny problem faced by researchers is how to dereplicate NPs from crude extracts. Remarkably, with the rapid development of omics, analytical instrumentation, and artificial intelligence technology, in 2012, an efficient approach, known as tandem mass spectrometry (MS/MS)-based molecular networking (MN) analysis, was developed to avoid the rediscovery of known compounds from the complex natural mixtures. Then, in the past decade, based on the classical MN (CLMN), feature-based MN (FBMN), ion identity MN (IIMN), building blocks-based molecular network (BBMN), substructure-based MN (MS2LDA), and bioactivity-based MN (BMN) methods have been presented. In this paper, we review the basic principles, general workflow, and application examples of the methods mentioned above, to further the research and applications of these methods.
Hey everyone!
Excited to be here and kind of a fresh start!
I'm an engineer and scientist that loves to build tools for the community. Primarily working on computational tools for the analysis of untargeted mass spectrometry data.
Very excited to start migrating and launching new tools at UC Riverside soon!




 ✡️ 🇵🇸
✡️ 🇵🇸