Thanks for a wonderful 1st day of the #IUMS2024. It's been busy at our booth yesterday 😎 Take your chance and visit us today to learn about the DSMZ 🦠🧫🧬🔬
Ulrich Nübel
- 130 Followers
- 167 Following
- 49 Posts
Very excited to announce our latest publication in Nucleic Acids Research!
https://doi.org/10.1093/nar/gkae902
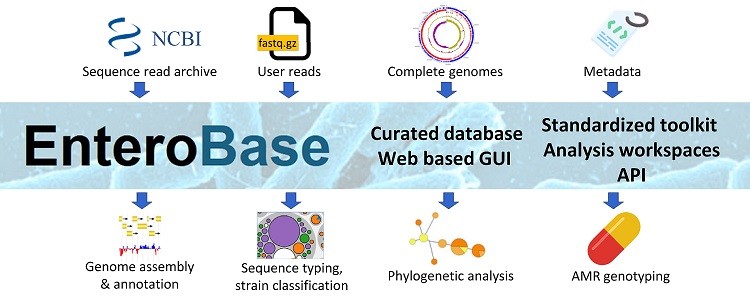
It provides an update on the content and tools of the EnteroBase platform for web-based pathogen genomic surveillance. Great international collaboration among #WarwickUniversity, #SoochowUniversity, @ForschungszentrumBorstel, @DrKatHolt and @dsmz, as part of #dzif and #pangens.
online petition in support of a proposal that is important for the future of prokaryote nomenclature:
https://www.ipetitions.com/petition/support-the-best-of-both-worlds
Petition Support the Best of Both Worlds
The SeqCode removes the incentive to cultivate and deposit from prokaryotic nomenclature, reduces scientific replicability, undermines a scientific standard, and is fundamentally unfair, giving equal status to names based on the accessible organism itself and names based solely on experimental results. The ICSP should not accept the SeqCode but ratify a more suitable alternative, the
The best of both worlds: a proposal for further integration of Candidatus names into the International Code of Nomenclature of Prokaryotes
The naming of prokaryotes is governed by the International Code of Nomenclature of Prokaryotes (ICNP) and partially by the International Code of Nomenclature for Algae, Fungi and Plants (ICN). Such codes must be able to determine names of taxa in a universal and unambiguous manner, thus serving as a common language across different fields and activities. This unity is undermined when a new code of nomenclature emerges that overlaps in scope with an established, time-tested code and uses the same format of names but assigns different nomenclatural status values to the names. The resulting nomenclatural confusion is not beneficial to the wider scientific community. Such ambiguity is expected to result from the establishment of the ‘Code of Nomenclature of Prokaryotes Described from DNA Sequence Data’ (‘SeqCode’), which is in general and specific conflict with the ICNP and the ICN. Shortcomings in the interpretation of the ICNP may have exacerbated the incompatibility between the codes. It is reiterated as to why proposals to accept sequences as nomenclatural types of species and subspecies with validly published names, now implemented in the SeqCode, have not been implemented by the International Committee on Systematics of Prokaryotes (ICSP), which oversees the ICNP. The absence of certain regulations from the ICNP for the naming of as yet uncultivated prokaryotes is an acceptable scientific argument, although it does not justify the establishment of a separate code. Moreover, the proposals rejected by the ICSP are unnecessary to adequately regulate the naming of uncultivated prokaryotes. To provide a better service to the wider scientific community, an alternative proposal to emend the ICNP is presented, which would result in Candidatus names being regulated analogously to validly published names. This proposal is fully consistent with previous ICSP decisions, preserves the essential unity of nomenclature and avoids the expected nomenclatural confusion.
Our paper on transcriptomics and metabolomics of the myxobacterium Sorangium sp. was published in Microbial Biotechnology today!
Great collaboration with @juanmobovi and @[email protected].
https://ami-journals.onlinelibrary.wiley.com/doi/epdf/10.1111/1751-7915.14246
Very excited that our paper is now published in Microbial Biotechnology (@MicrobialBiote1
)! Finding the time point of BGC transcription and #NaturalProduct production in a #myxobacteria by linking #genomics, #transcriptomics, and #metabolomics
🐦 🔗 https://twitter.com/juanmobovi/status/1641776788312293376
Judith Boldt on Twitter
“Very excited that our paper is now published in Microbial Biotechnology (@MicrobialBiote1)! Finding the time point of BGC transcription and #NaturalProduct production in a #myxobacteria by linking #genomics, #transcriptomics, and #metabolomics. https://t.co/3w1JUZvzIB”
We're so happy to have you all here on mstdn.science! We want this instance to be transparent in all the best ways, including how we spend donations. 💰
So, we've teamed up with Open Collective and now have an account where you can donate directly - and see how we spend the money! (Only on the running costs: servers, storage, etc 💻 )
Check it out, and become a contributor at https://opencollective.com/mstdnscience !
Even a few $/£/€ a month absolutely make a difference - and thank you for helping us!! 🙏 😁
gutSMASH predicts specialized primary metabolic pathways from the human gut microbiota
Four principles to establish a universal virus taxonomy
Transforming an existing phenotypic classification of viruses into one based on evolutionary relationships that can accommodate the vast number of viruses characterized in metagenomics and environmental studies is an ongoing challenge. This Consensus View explains how such a taxonomy can be expanded to encapsulate viral diversity and to recognize independent biological origins of different virus groups.
I am happy to share the news that our latest paper was published in Environmental Microbiology:
https://ami-journals.onlinelibrary.wiley.com/doi/epdf/10.1111/1462-2920.16352
Here we report the usage of bacterial genome sequencing for tracing the insect-associated spread of bacterial pathogens back to their source in a pig farm.
Great collaboration with ATB, ZALF (@ZALF_leibniz), and LVAT within the Leibniz Research Alliance INFECTIONS (https://leibnizinfections.de/en/).