Thrilled to announce that our preprint (together with fantastic @richardneher and Liam Shaw) has been officially published in Molecular Biology and Evolution! 🎉
If you're curious about how rapidly the genome of E. coli evolves (picking up new genes, rearranging, breaking synteny...) give it a look! 🧬
https://academic.oup.com/mbe/advance-article/doi/10.1093/molbev/msae272/7942412
A summary thread 🧵 https://mstdn.science/@mrcmlr/112772499102225806
[2/13]
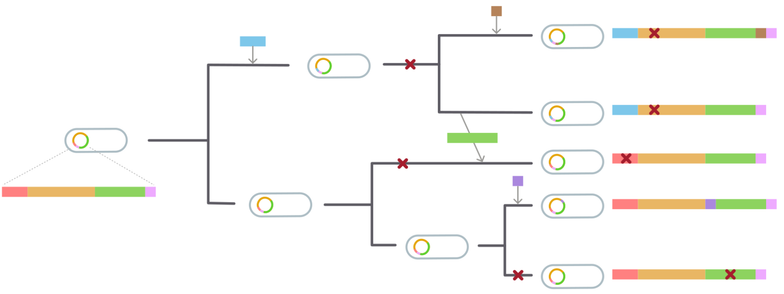
Horizontal Gene Transfer, i.e. the exchange of genetic material from one individual to the next, is at the origin of many of these changes. This process however also complicates phylogenetic inference, invalidating the hypothesis of exclusively vertical inheritance.
Happy to announce a new preprint from the lab, together with fantastic @richardneher and Liam Shaw! 📝
https://www.biorxiv.org/content/10.1101/2024.07.08.602537v1
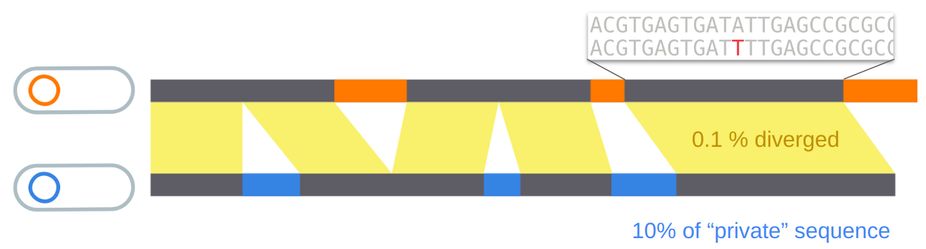
We were motivated by the following puzzling observation: microbial genomes can be extremely similar in the core genome, while still differing by large portions of accessory genome.
Therefore we ask the question:
1) how does this diversity accumulate?
2) at what rate do genomes undergo large structural changes?
A thread 🧵 [1/13]