#CompChem #CompBiol #AI #Biotech
lab_colombo
- 123 Followers
- 128 Following
- 171 Posts
#CompChem #CompBiol #AI #Biotech
https://pubs.acs.org/doi/full/10.1021/jacs.3c11396
https://pubs.acs.org/doi/10.1021/acs.jmedchem.3c01236
Hi Structural biologists!
Here is our article that helps you make the case that the experiment is still very much needed: https://www.nature.com/articles/s41592-023-02087-4
"AlphaFold predictions are valuable hypotheses and accelerate but do not replace experimental structure determination." Nature Methods (2023)
#Crystallography #MX #PDB #Alphafold #StructuralBiology @strucbio
AlphaFold predictions are valuable hypotheses and accelerate but do not replace experimental structure determination - Nature Methods
An analysis of AlphaFold protein structure predictions shows that while in many cases the predictions are highly accurate, there are also many instances where the predicted structures or parts of predicted structures do not agree with experimentally resolved data. Therefore, care must be taken when using these predictions for informing structural hypotheses.
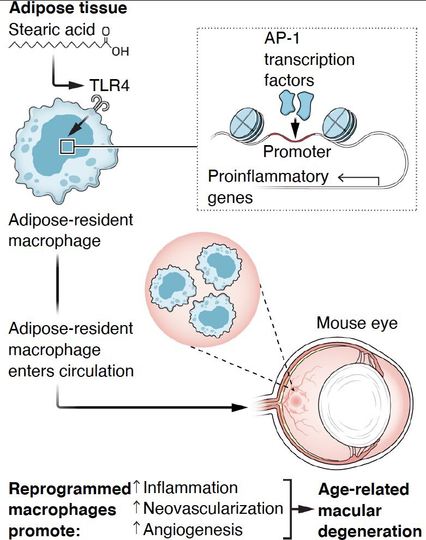
Obesity confers #macrophage memory
◽️Obesity triggers #epigenetic reprogramming of the innate immunity, lasting long after return to normal weight
◽️Stearic fatty acid (via TLR4) remodels chromatin & ↑AP-1 accessibility
◽️MΦ shift OXPHOS→ glycolysis
RT (X) |
https://twitter.com/stshuken/status/1711399515410809110
---
#proteomics #prot-RT #prot-paper
Steven Shuken on X
My first @GygiLab paper is out in @an_chem! Under the mentorship of postdoc @qingyuhms I profiled electrophilic compounds’ effects on the proteome at >8k protein depth. The new Ascend takes 30% less time, improves quant, reveals relevant scarce proteins. https://t.co/LrBOAGnNl8