Do you (want to) use whole-plasmid sequencing (e.g. @plasmidsaurus) but are put off by $15/sample? Use SAVEMONEY—our (postdoc Masaaki Uematsu’s) algorithm that lets you mix 6 or more plasmids/sample & computationally de-mixes reads!
https://www.biorxiv.org/content/10.1101/2023.04.12.536413v1
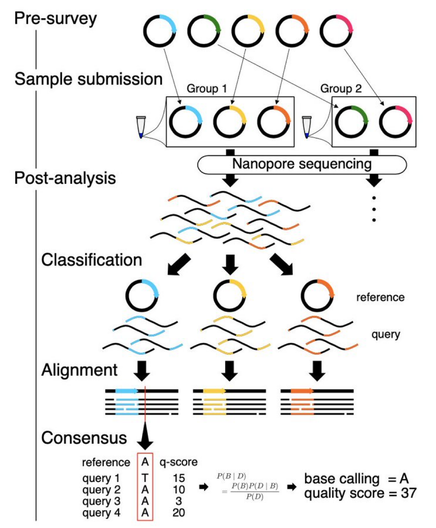
How it works: 1/n